Details of Host Protein
| Host Protein General Information (ID: PT0258) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
CCR4-associated factor 4 (NOT4)
|
Gene Name |
CNOT4
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
CNOT4_HUMAN
|
||||||
| EC Number |
2.3.2.27
|
||||||||
| Subcellular Location |
Cytoplasm Nucleus
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Has E3 ubiquitin ligase activity, promoting ubiquitinationand degradation of target proteins. Involved in activation of the JAK/STAT pathway. Catalyzes ubiquitination ofmethylated RBM15. Plays a role in quality control oftranslation of mitochondrial outer membrane-localized mRNA. As part of the PINK1-regulated signaling, uponmitochondria damage, ubiquitinates ABCE1 and thereby recruits autophagyreceptors to the mitochondrial outer membrane to initiate mitophagy.
[1-4]
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| RNA degradation | hsa03018 |
Pathway Map 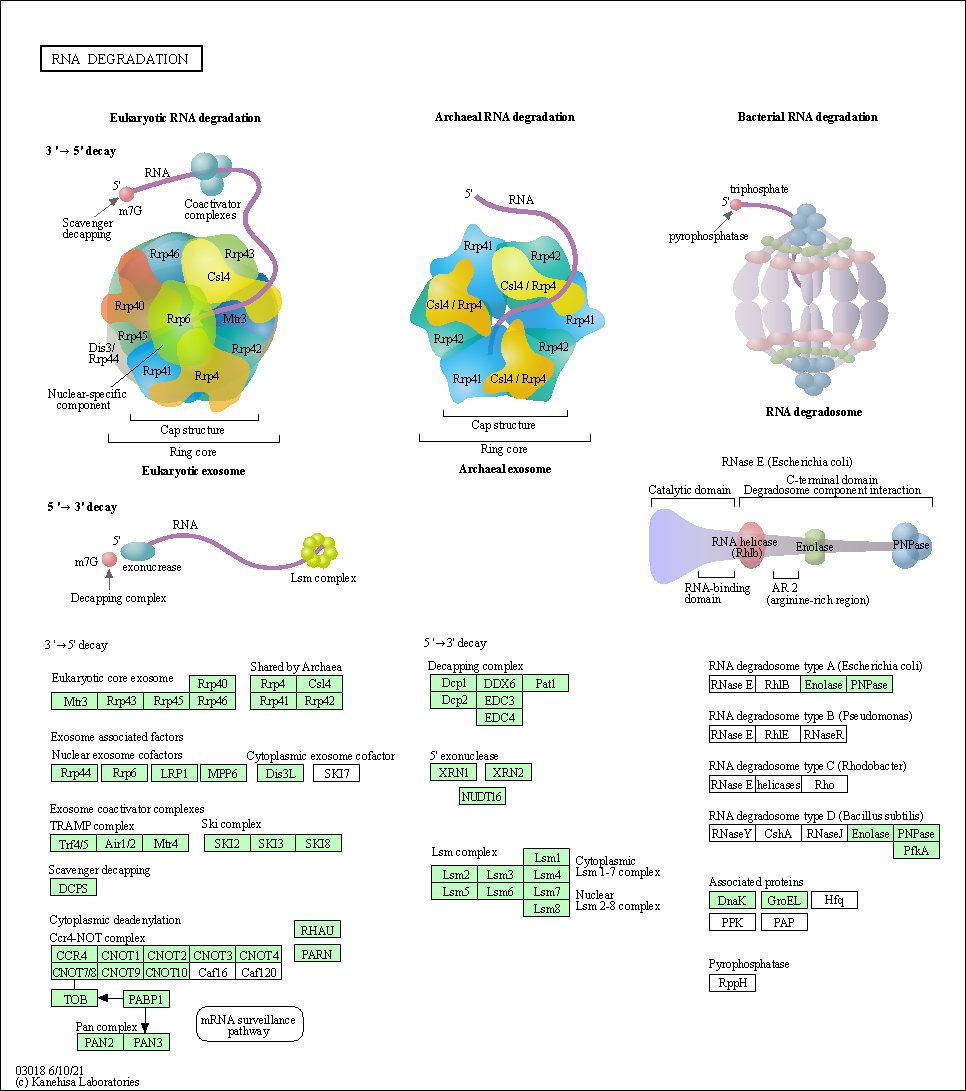
|
|||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Anti-viral | [5] | |||||
| Infected Tissue | Lung | Infection Time | 36 h | ||||||
| Infected Cell | A549 Cells (Adenocarcinomic Human alveolar basal epithelial cells) | Cellosaurus ID | CVCL_H249 | ||||||
| Method Description | To detect the role of host protein CNOT4 in viral infection, CNOT4 protein knockout A549 Cells were infected with SARS-COV-2 for 36 h , and the effects on infection was detected through WB. | ||||||||
| Results | It is reported that Knockdown of CNOT4 increases infection compared with control group. | ||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: ORF1ab (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF7a (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF7a
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF7b (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF7b
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: N region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: S region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
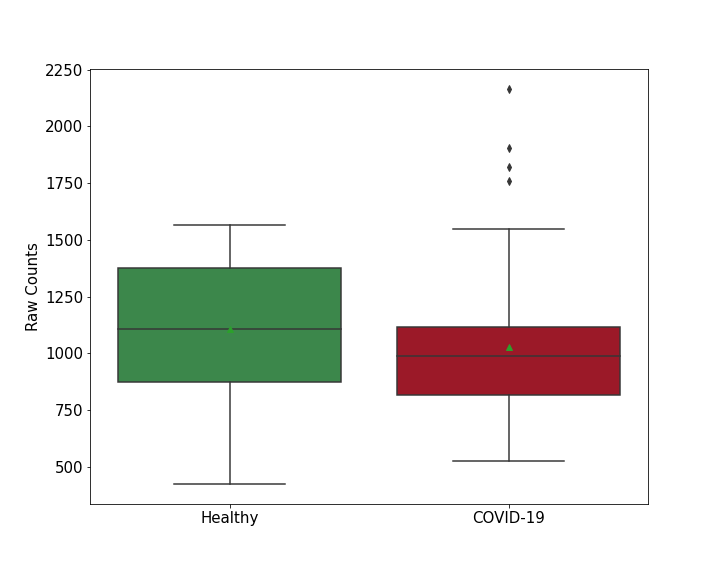
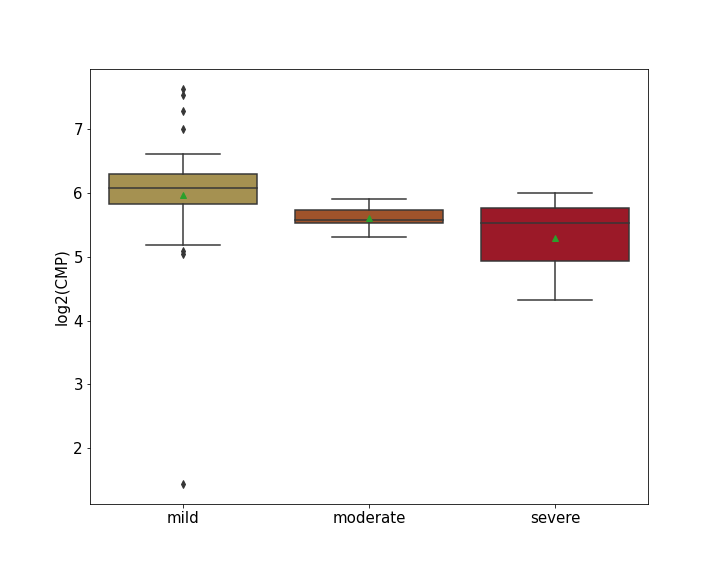
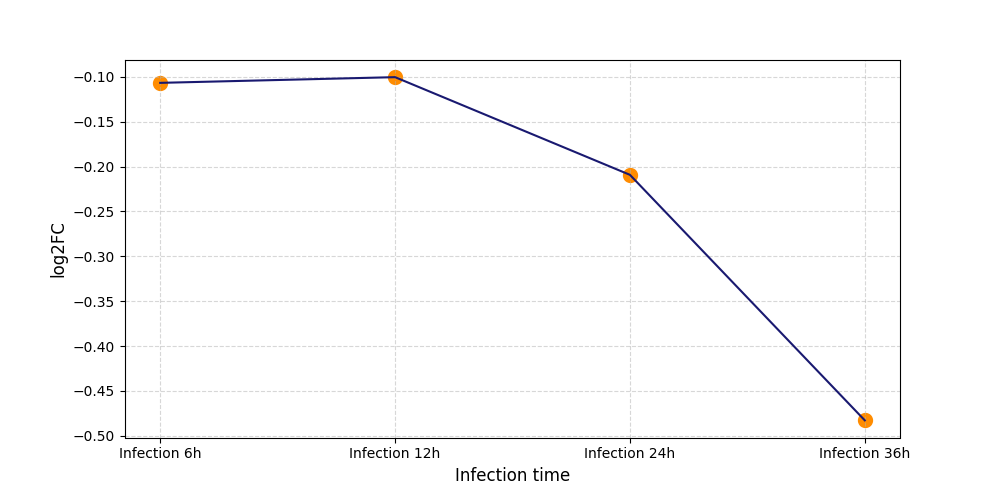
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
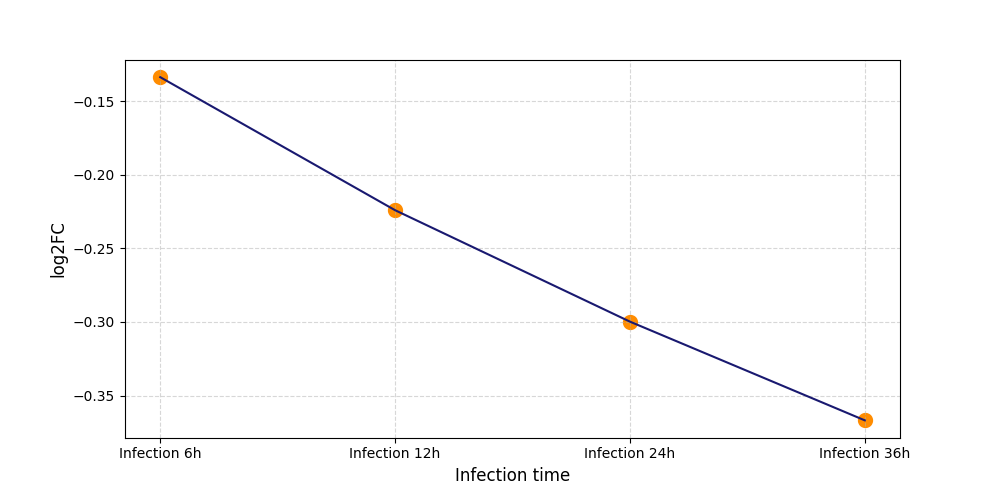
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S301
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
S324
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
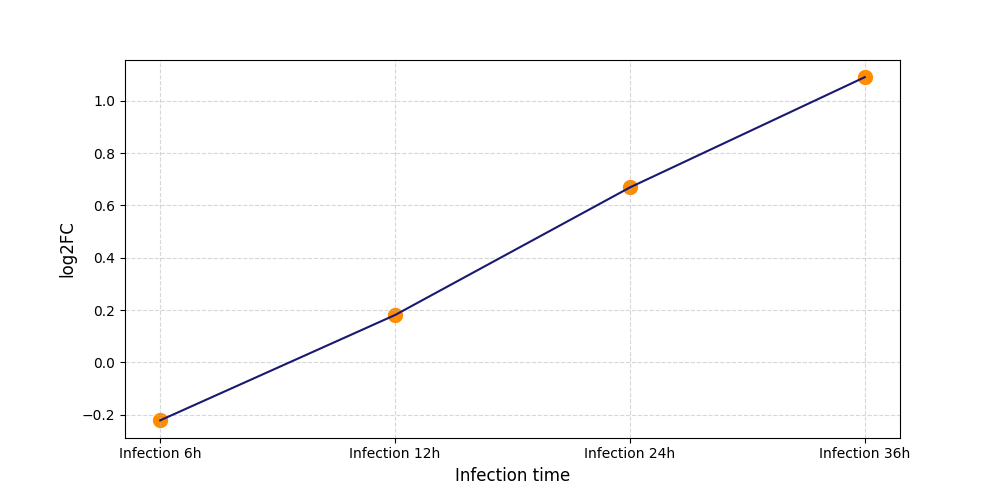
S432
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MSRSPDAKEDPVECPLCMEPLEIDDINFFPCTCGYQICRFCWHRIRTDENGLCPACRKPYPEDPAVYKPLSQEELQRIKNEKKQKQNERKQKISENRKHLASVRVVQKNLVFVVGLSQRLADPEVLKRPEYFGKFGKIHKVVINNSTSYAGSQGPSASAYVTYIRSEDALRAIQCVNNVVVDGRTLKASLGTTKYCSYFLKNMQCPKPDCMYLHELGDEAASFTKEEMQAGKHQEYEQKLLQELYKLNPNFLQLSTGSVDKNKNKVTPLQRYDTPIDKPSDSLSIGNGDNSQQISNSDTPSPPPGLSKSNPVIPISSSNHSARSPFEGAVTESQSLFSDNFRHPNPIPSGLPPFPSSPQTSSDWPTAPEPQSLFTSETIPVSSSTDWQAAFGFGSSKQPEDDLGFDPFDVTRKALADLIEKELSVQDQPSLSPTSLQNSSSHTTTAKGPGSGFLHPAAATNANSLNSTFSVLPQRFPQFQQHRAVYNSFSFPGQAARYPWMAFPRNSIMHLNHTANPTSNSNFLDLNLPPQHNTGLGGIPVAGEEEVKVSTMPLSTSSHSLQQGQQPTSLHTTVA
Click to Show/Hide
|
|---|