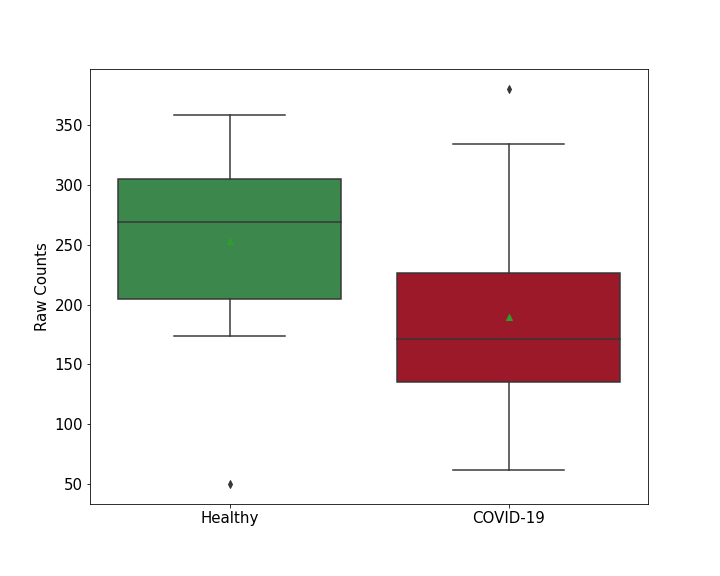
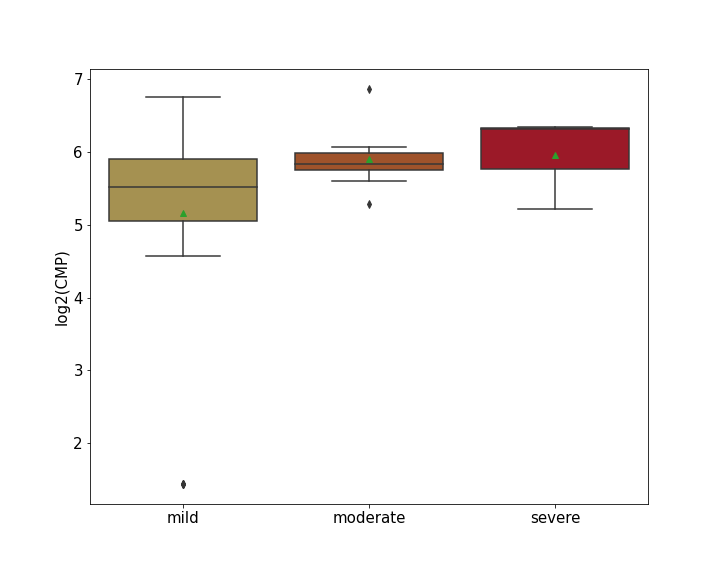
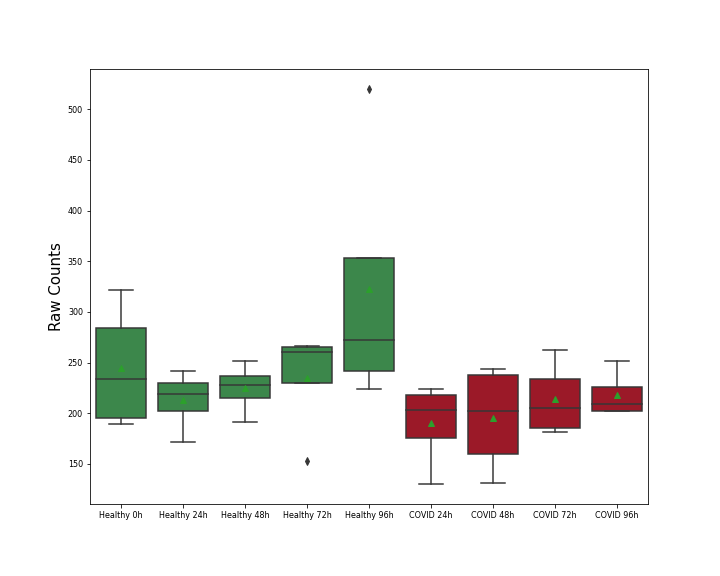
Details of Host Protein
| Host Protein General Information (ID: PT0207) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
DEAD box protein 19A (DDX19L)
|
Gene Name |
DDX19A
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
DD19A_HUMAN
|
||||||
| Protein Families |
DEAD box helicase family
|
||||||||
| EC Number |
3.6.4.13
|
||||||||
| Subcellular Location |
Cytoplasm Nucleus; nucleoplasm
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
ATP-dependent RNA helicase involved in mRNA export from thenucleus. Rather than unwinding RNA duplexes, DDX19 functions as aremodeler of ribonucleoprotein particles, whereby proteins bound tonuclear mRNA are dissociated and replaced by cytoplasmic mRNA bindingproteins.
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| Nucleocytoplasmic transport | hsa03013 |
Pathway Map 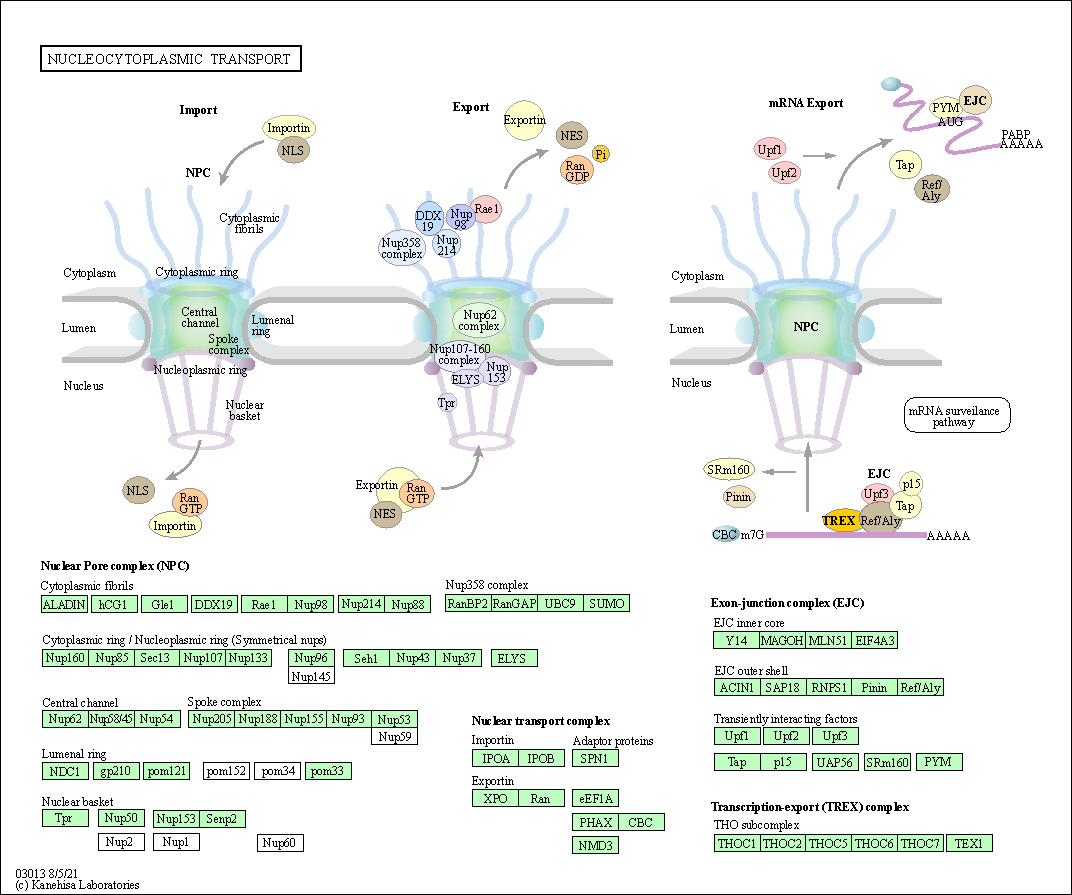
|
|||||||
| mRNA surveillance pathway | hsa03015 |
Pathway Map 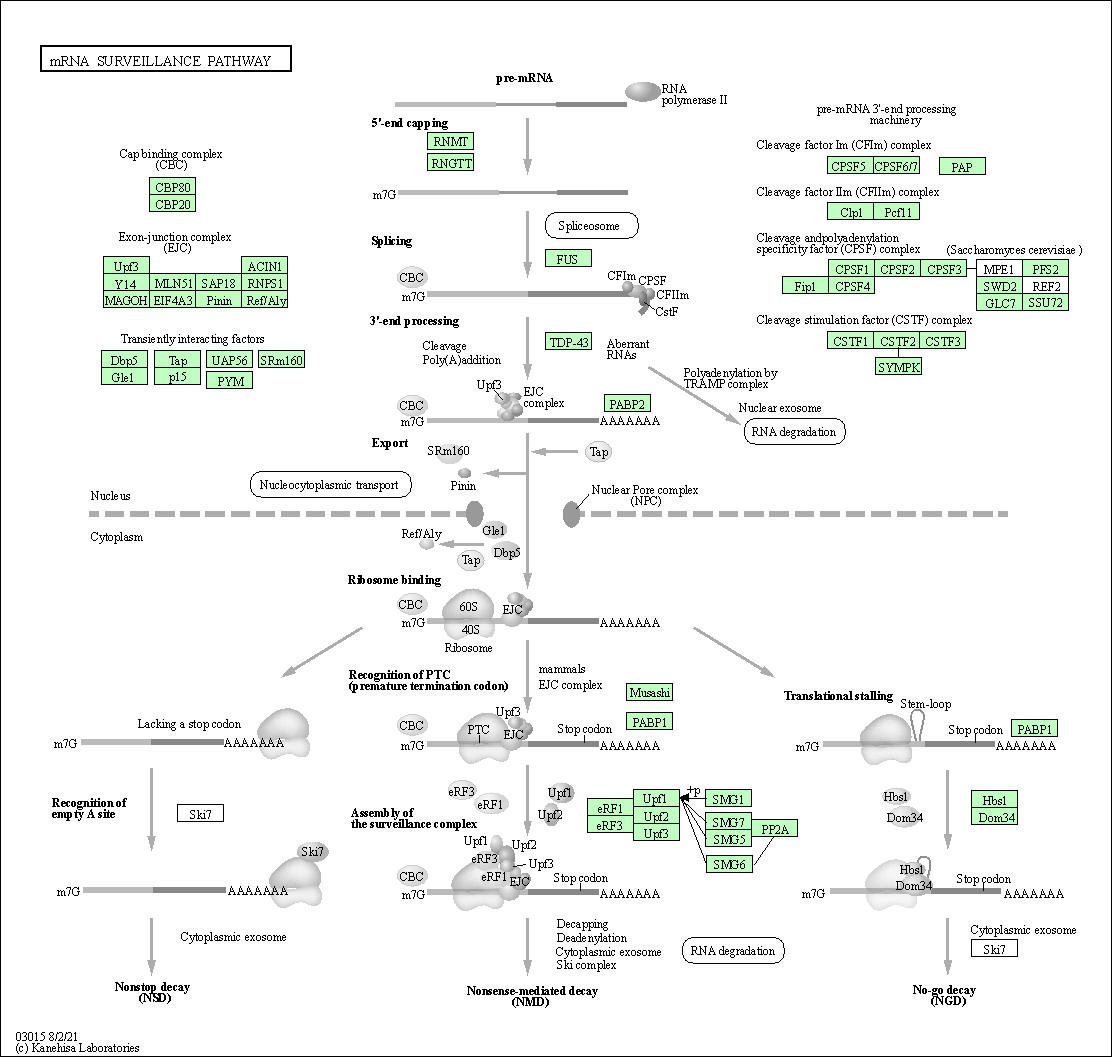
|
|||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Anti-viral | [1] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein DDX19A in viral infection, DDX19A protein knockout Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of DDX19A increases SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: Not Specified Virus Region (hCoV-19/USA/WA1/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[2] | |||||||
| Strains Name |
hCoV-19/USA/WA1/2020
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7.5 cells (Hepatocyte derived cellular carcinoma cell) (CVCL_7927 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 48 h | ||||||||
| Interaction Score | FDR ≤ 0.05 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||