Details of Host Protein
| Host Protein General Information (ID: PT0430) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
Exosome component 10 (POLR2A)
|
Gene Name |
EXOSC10
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
EXOSX_HUMAN
|
||||||
| EC Number |
3.1.13.-
|
||||||||
| Subcellular Location |
Cytoplasm
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Putative catalytic component of the RNA exosome complex whichhas 3'->5' exoribonuclease activity and participates in a multitude ofcellular RNA processing and degradation events. In the nucleus, the RNAexosome complex is involved in proper maturation of stable RNA speciessuch as rRNA, snRNA and snoRNA, in the elimination of RNA processingby-products and non-coding 'pervasive' transcripts, such as antisenseRNA species and promoter-upstream transcripts (PROMPTs), and of mRNAswith processing defects, thereby limiting or excluding their export tothe cytoplasm. The RNA exosome may be involved in Ig class switchrecombination (CSR) and/or Ig variable region somatic hypermutation (SHM) by targeting AICDA deamination activity to transcribed dsDNAsubstrates. In the cytoplasm, the RNA exosome complex is involved ingeneral mRNA turnover and specifically degrades inherently unstablemRNAs containing AU-rich elements (AREs) within their 3' untranslatedregions, and in RNA surveillance pathways, preventing translation ofaberrant mRNAs. It seems to be involved in degradation of histone mRNA. EXOSC10 has 3'-5' exonuclease activity. EXOSC10 isrequired for nucleolar localization of C1D and probably mediates theassociation of MTREX, C1D and MPHOSPH6 with the RNA exosome involved inthe maturation of 5. 8S rRNA.
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| RNA degradation | hsa03018 |
Pathway Map 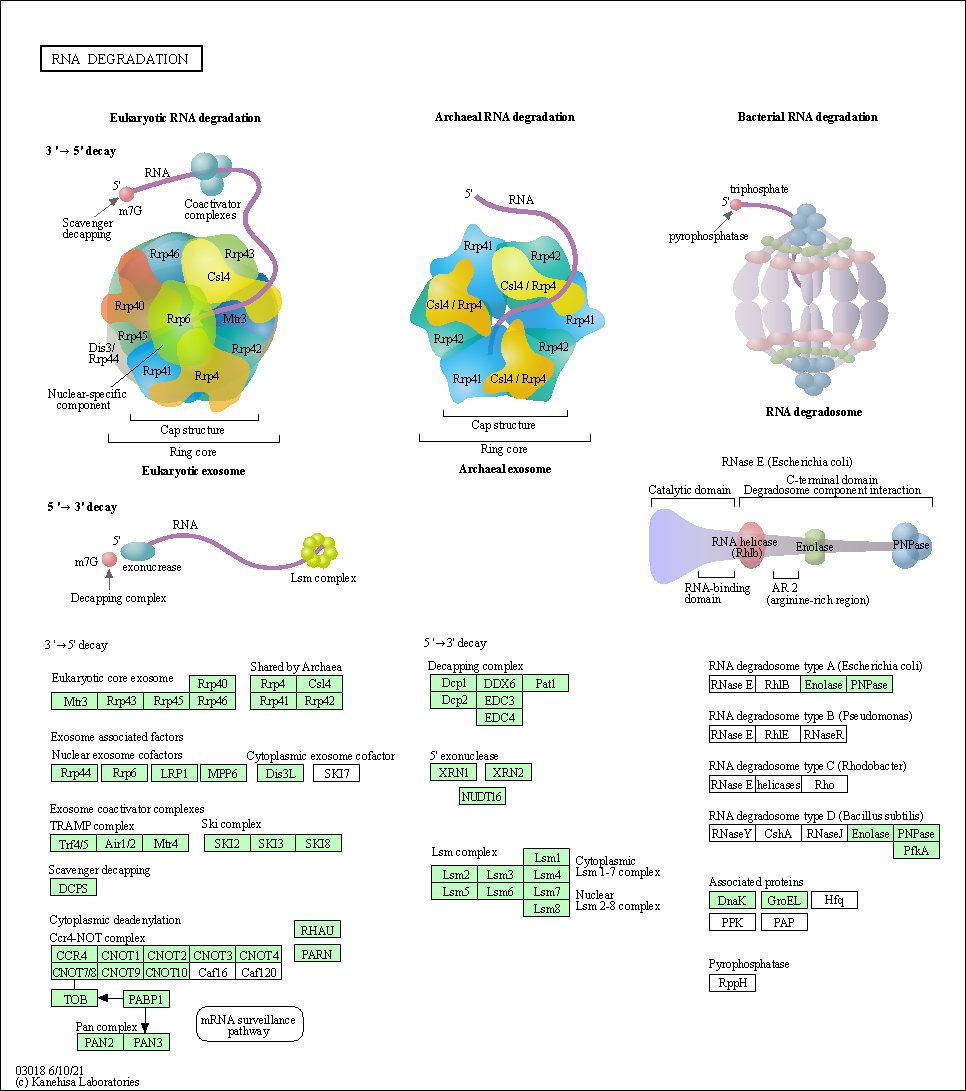
|
|||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Pro-viral | [1] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein EXOSC10 in viral infection, EXOSC10 protein knockout Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of EXOSC10 leads to the decreased SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: ORF10 (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[2] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
ORF10
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Calu-3 cells (Human Lung Cancer Cell) (CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||