Details of Host Protein
| Host Protein General Information (ID: PT0502) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
Heat shock 70 kDa protein 1-like (ARHGAP23)
|
Gene Name |
HSPA1L
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
HS71L_HUMAN
|
||||||
| Protein Families |
Heat shock protein 7. family
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Molecular chaperone implicated in a wide variety of cellularprocesses, including protection of the proteome from stress, foldingand transport of newly synthesized polypeptides, activation ofproteolysis of misfolded proteins and the formation and dissociation ofprotein complexes. Plays a pivotal role in the protein quality controlsystem, ensuring the correct folding of proteins, the re-folding ofmisfolded proteins and controlling the targeting of proteins forsubsequent degradation. This is achieved through cycles of ATP binding, ATP hydrolysis and ADP release, mediated by co-chaperones. The affinityfor polypeptides is regulated by its nucleotide bound state. In theATP-bound form, it has a low affinity for substrate proteins. However, upon hydrolysis of the ATP to ADP, it undergoes a conformational changethat increases its affinity for substrate proteins. It goes throughrepeated cycles of ATP hydrolysis and nucleotide exchange, whichpermits cycles of substrate binding and release. Positive regulator of PRKN translocation to damaged mitochondria.
[1-2]
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| Antigen processing and presentation | hsa04612 |
Pathway Map 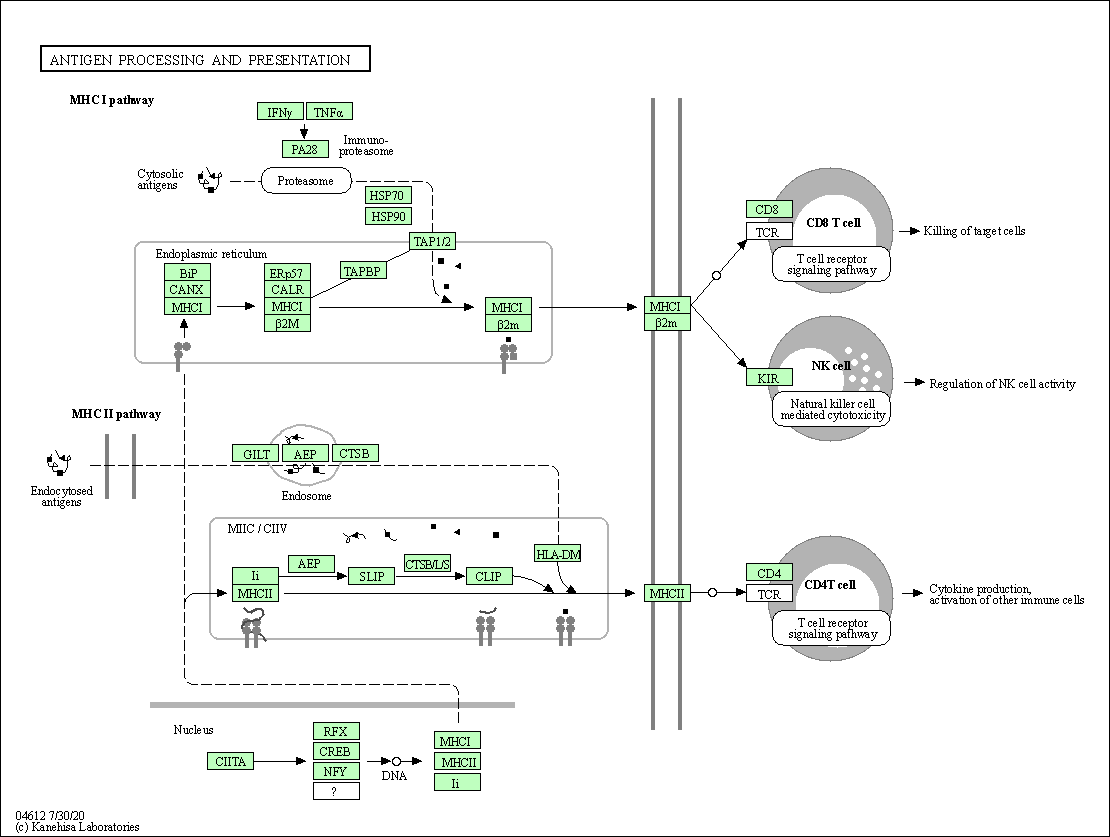
|
|||||||
| Measles | hsa05162 |
Pathway Map 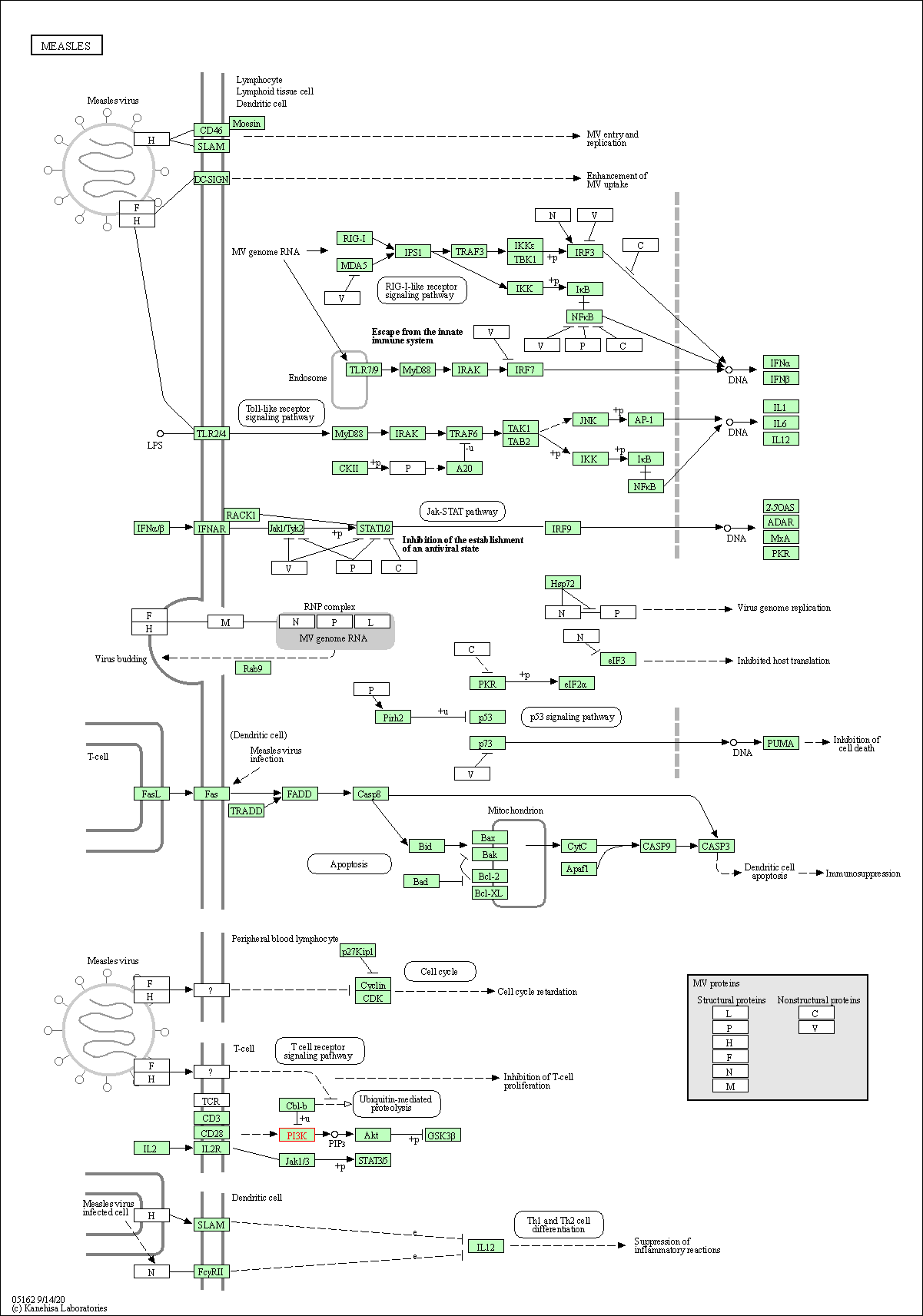
|
|||||||
| Legionellosis | hsa05134 |
Pathway Map 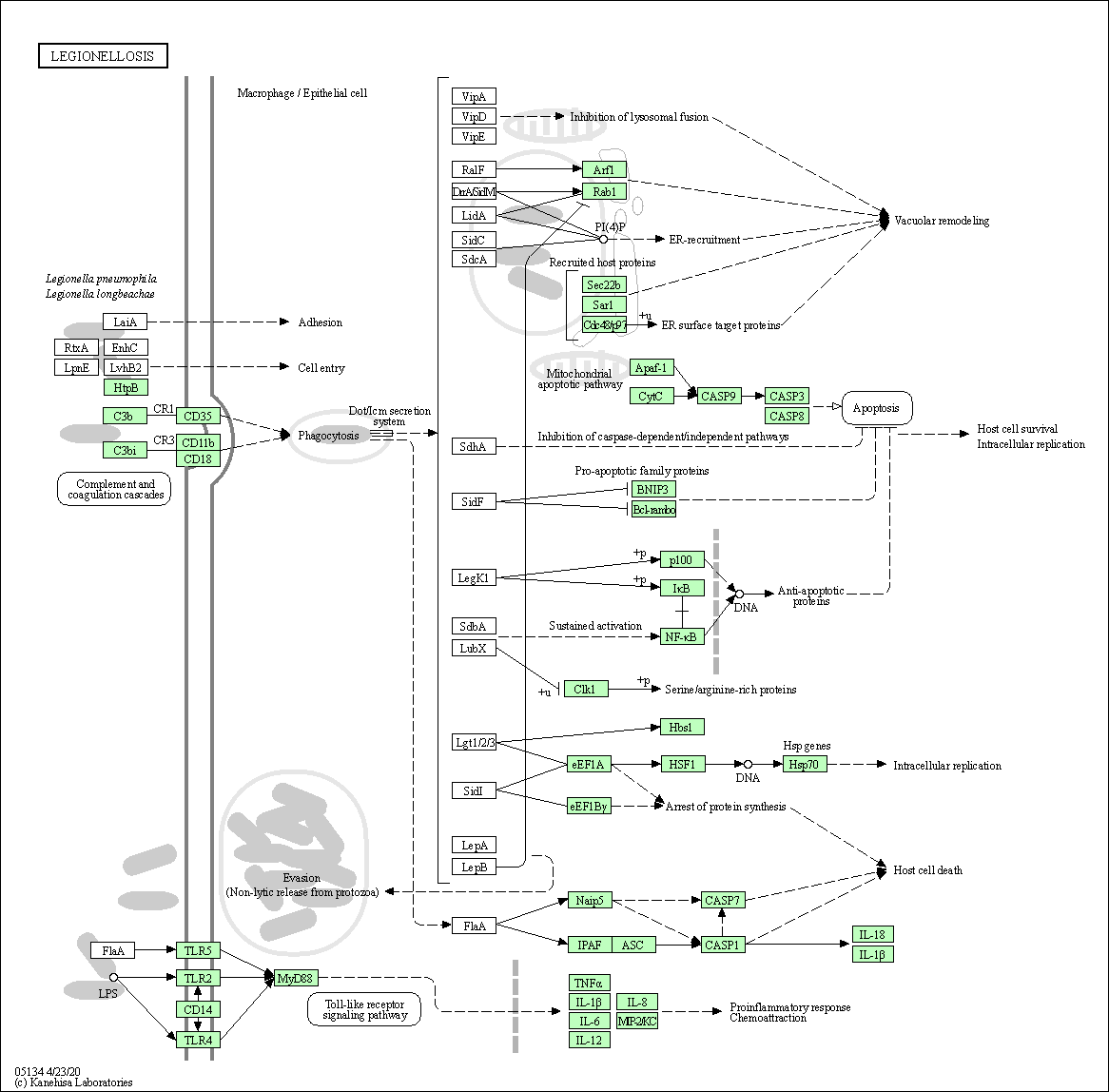
|
|||||||
| Toxoplasmosis | hsa05145 |
Pathway Map 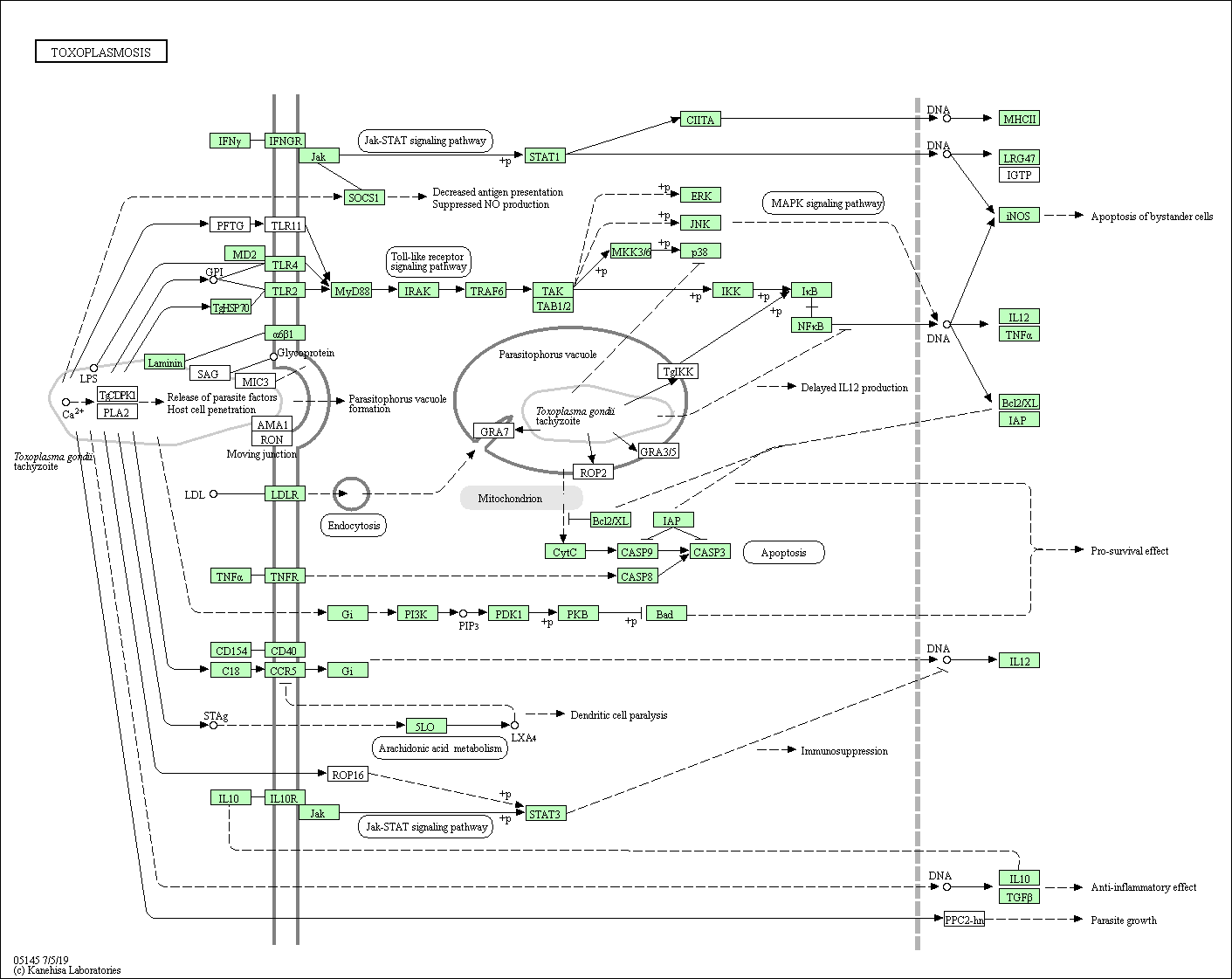
|
|||||||
| Protein processing in endoplasmic reticulum | hsa04141 |
Pathway Map 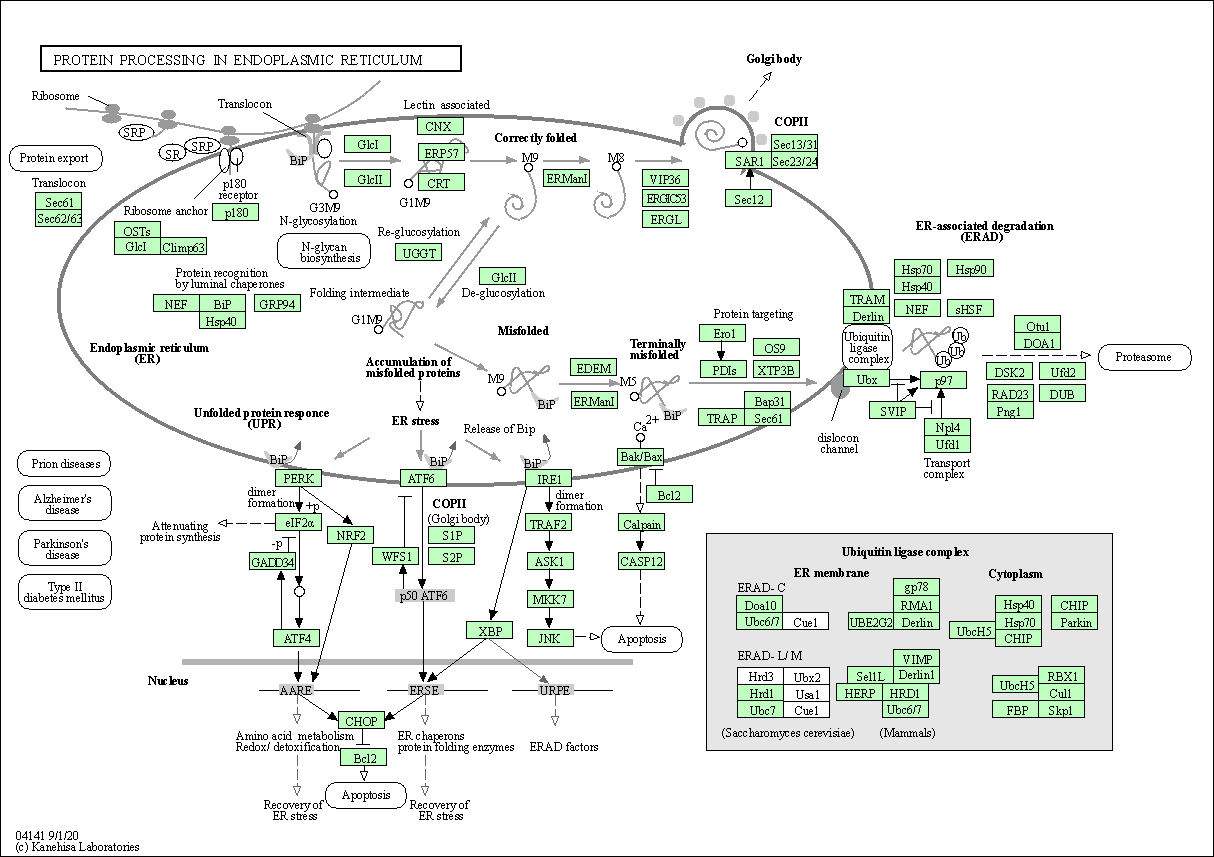
|
|||||||
| Spliceosome | hsa03040 |
Pathway Map 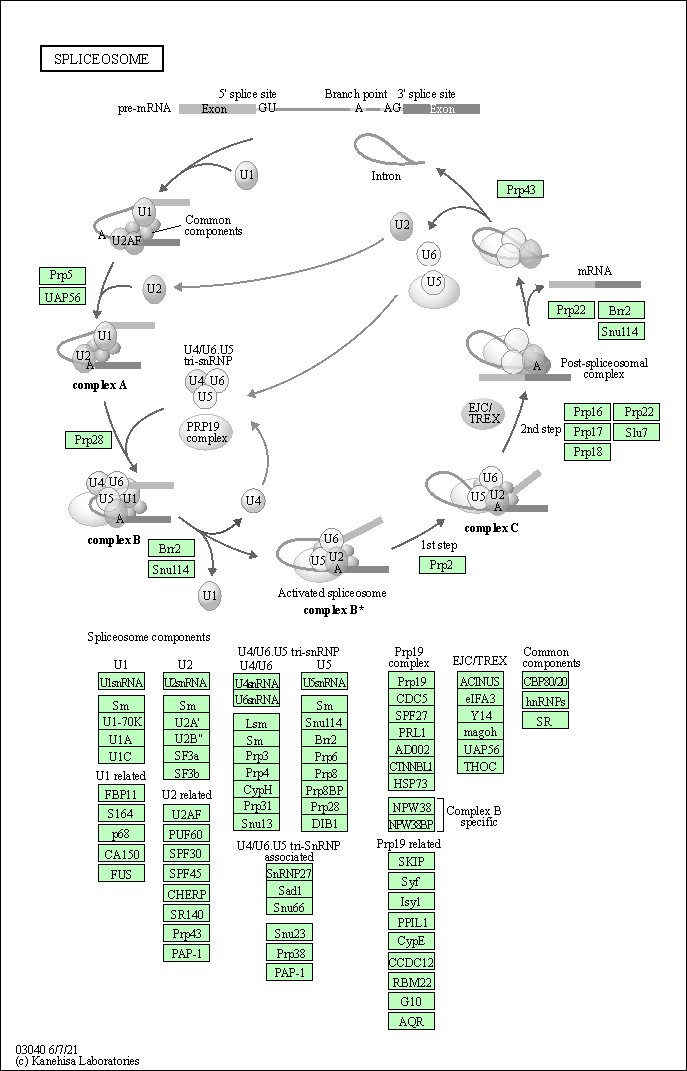
|
|||||||
| MAPK signaling pathway | hsa04010 |
Pathway Map 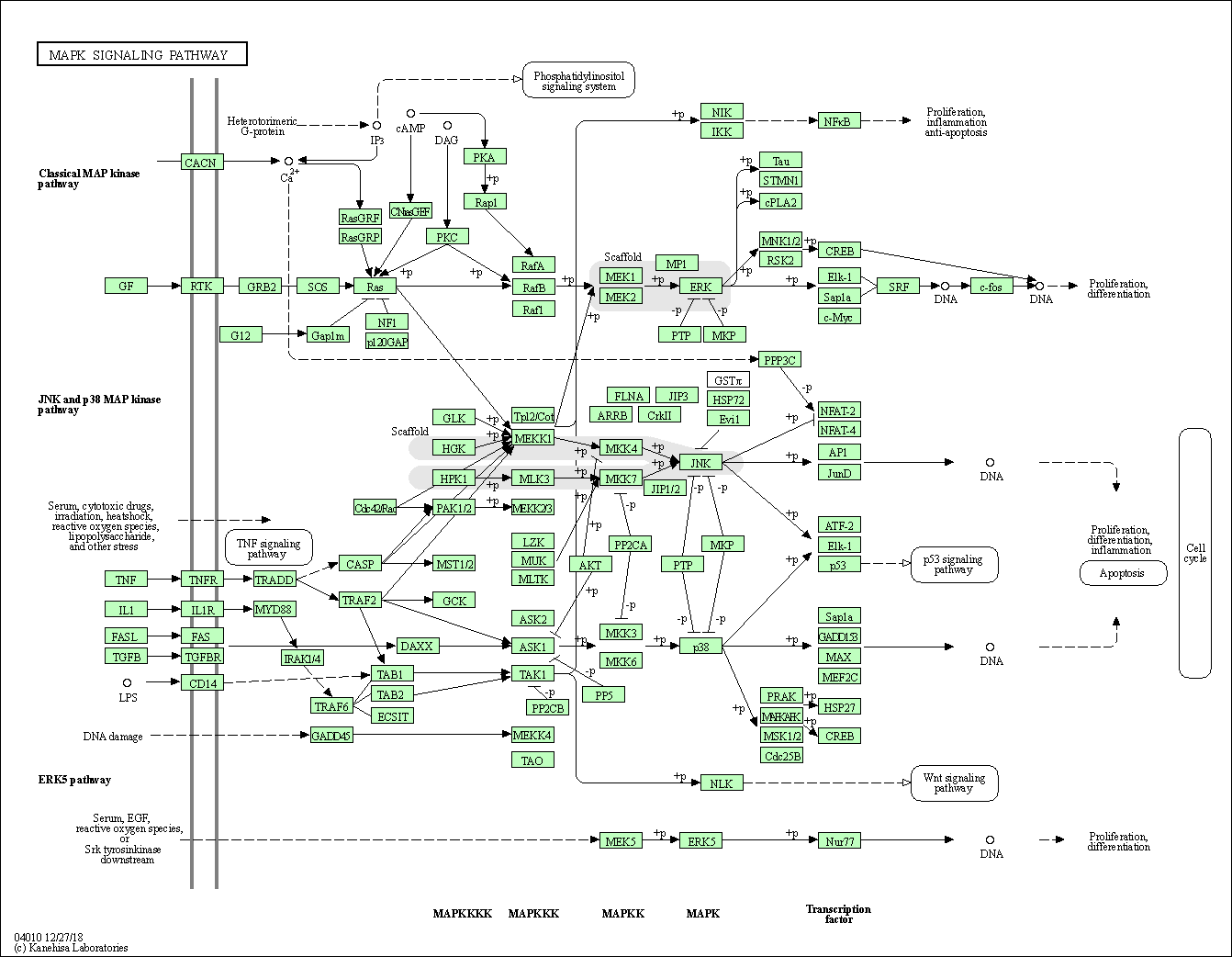
|
|||||||
| Endocytosis | hsa04144 |
Pathway Map 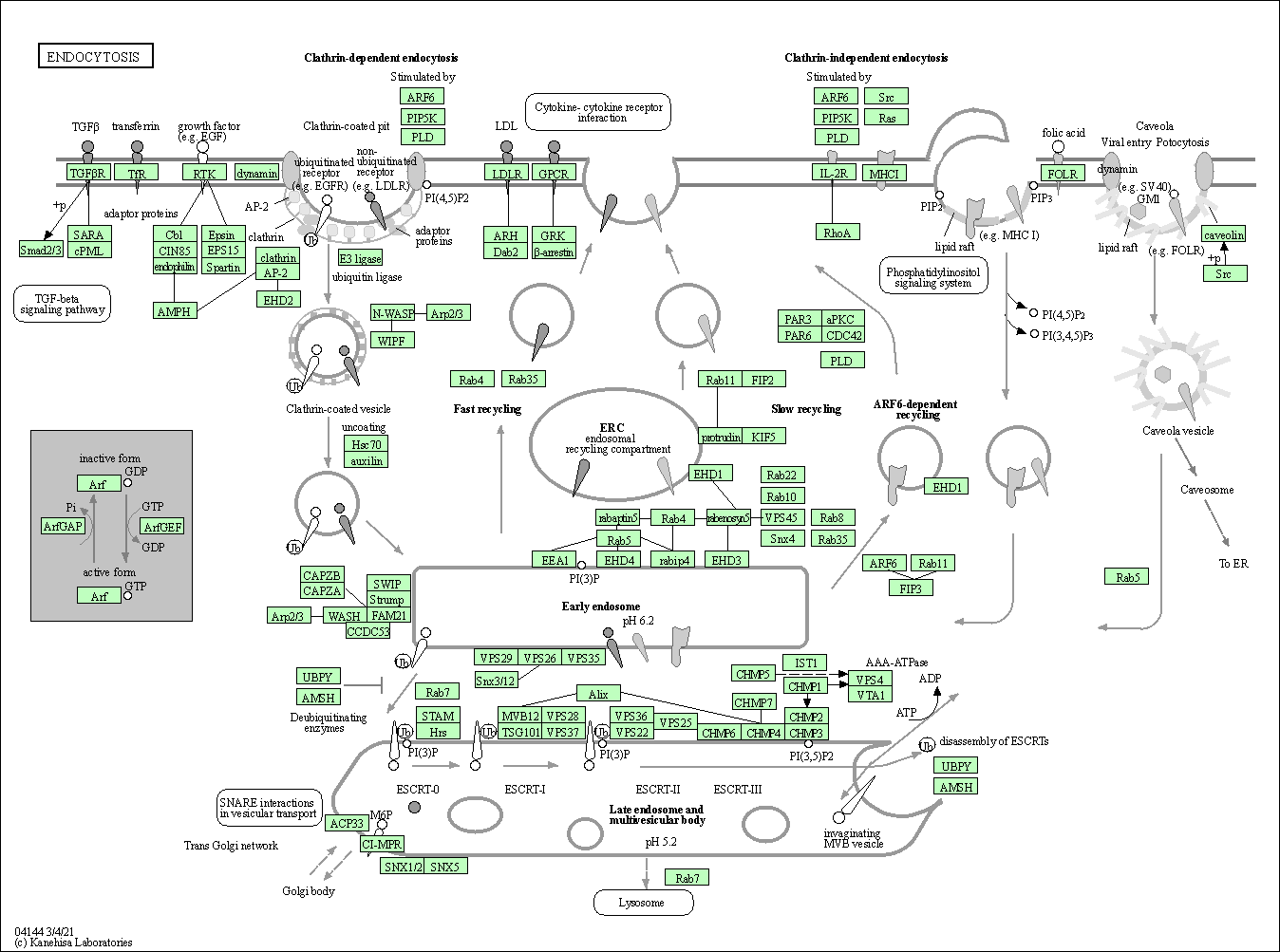
|
|||||||
| Longevity regulating pathway - multiple species | hsa04213 |
Pathway Map 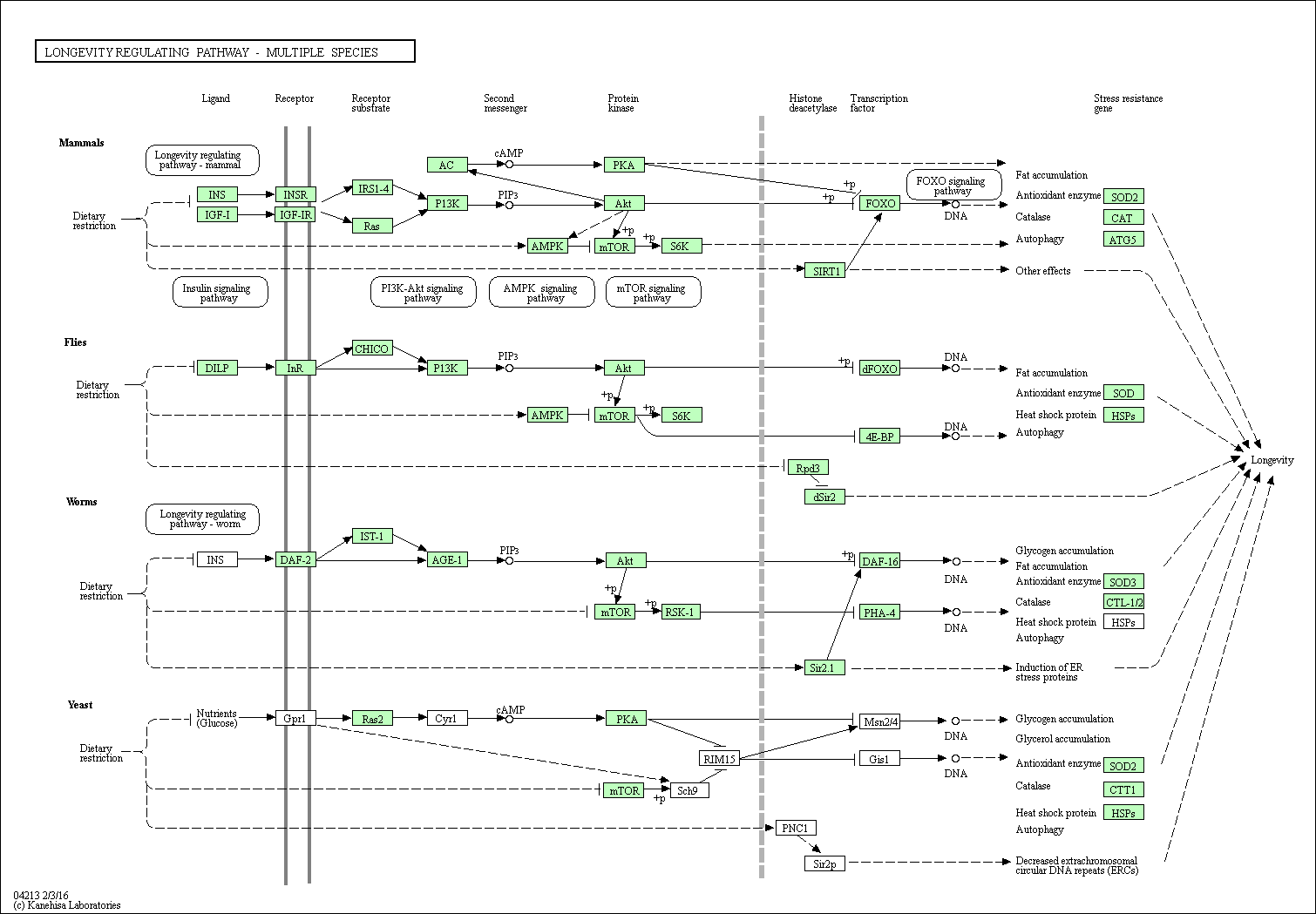
|
|||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Pro-viral | [3] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein HSPA1L in viral infection, HSPA1L protein knockout Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of HSPA1L leads to the decreased SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: 5'-UTR (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[4] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | HEK293 Cells (Human embryonic kidney cell) (CVCL_0045 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 48 h | ||||||||
| Interaction Score | Prot score = 492 | ||||||||
| Method Description | RNA-protein interaction detection (RaPID) assay; liquid chromatography with tandem mass spectrometry (LC-MS/MS) | ||||||||