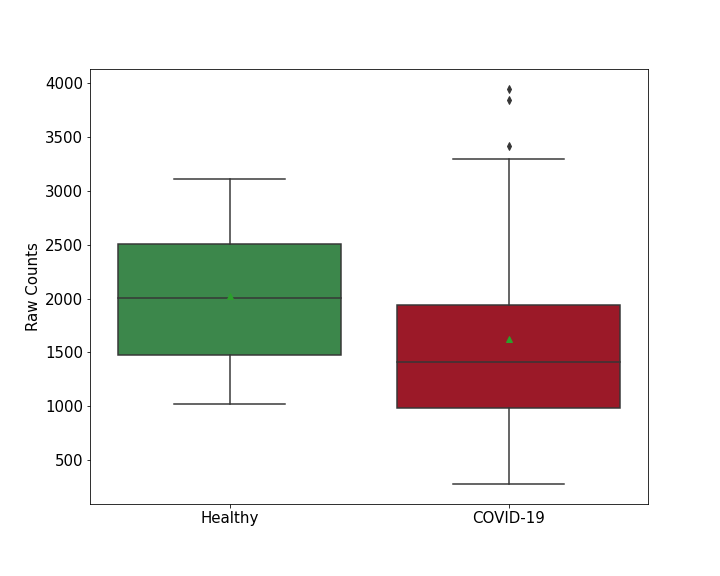
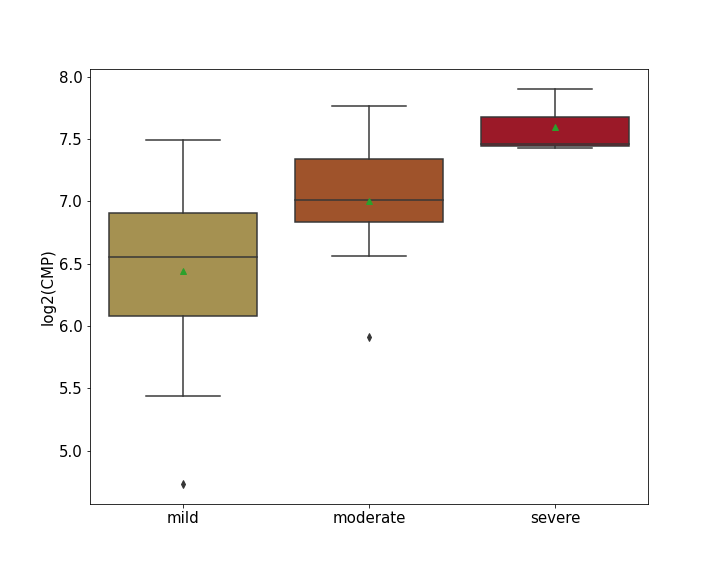
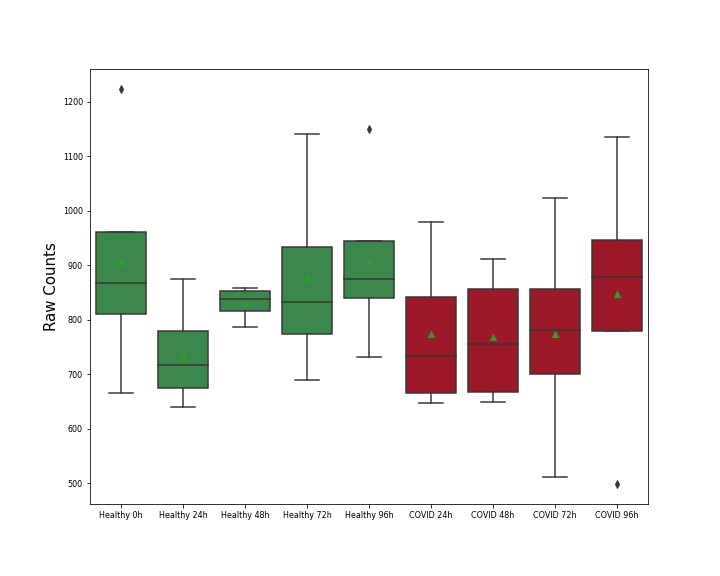
Details of Host Protein
| Host Protein General Information (ID: PT0558) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
Host cell factor 1 (HCFC1)
|
Gene Name |
HCFC1
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
HCFC1_HUMAN
|
||||||
| Subcellular Location |
Cytoplasm Nucleus
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Transcriptional coregulator. Involved incontrol of the cell cycle. Also antagonizestransactivation by ZBTB17 and GABP2; represses ZBTB17 activation of thep15 (INK4b) promoter and inhibits its ability to recruit p300. Coactivator for EGR2 and GABP2. Tethers the chromatin modifyingSet1/Ash2 histone H3 'Lys-4' methyltransferase (H3K4me) and Sin3histone deacetylase (HDAC) complexes (involved in the activation andrepression of transcription, respectively) together. Component of a THAP1/THAP3-HCFC1-OGT complex that is required for theregulation of the transcriptional activity of RRM1. As part of the NSL complex it may be involved in acetylation ofnucleosomal histone H4 on several lysine residues. Recruits KMT2E/MLL5 to E2F1 responsive promoters promotingtranscriptional activation and thereby facilitates G1 to S phasetransition. Modulates expression of homeobox proteinPDX1, perhaps acting in concert with transcription factor E2F1, therebyregulating pancreatic beta-cell growth and glucose-stimulated insulinsecretion. May negatively modulate transcriptionalactivity of FOXO3.
[1-4]
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| Herpes simplex virus 1 infection | hsa05168 |
Pathway Map 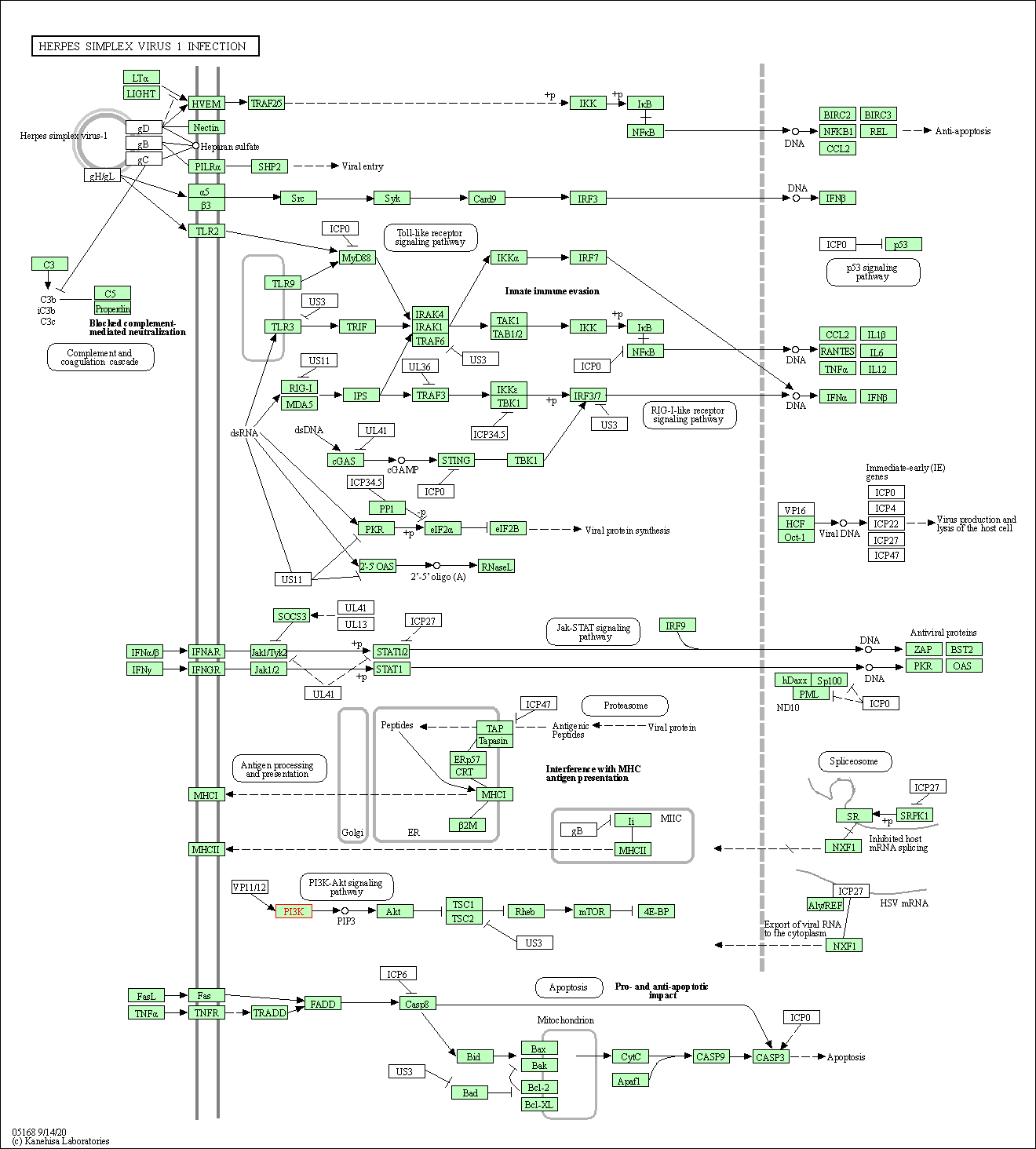
|
|||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Pro-viral | [5] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein HCFC1 in viral infection, HCFC1 protein knockout Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of HCFC1 leads to the decreased SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: ORF10 (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
ORF10
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Calu-3 cells (Human Lung Cancer Cell) (CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
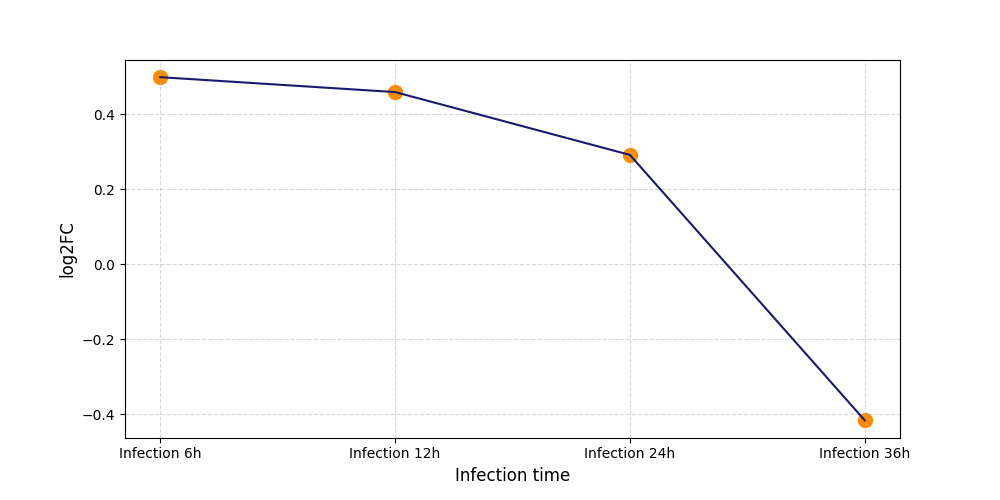
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S1205
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
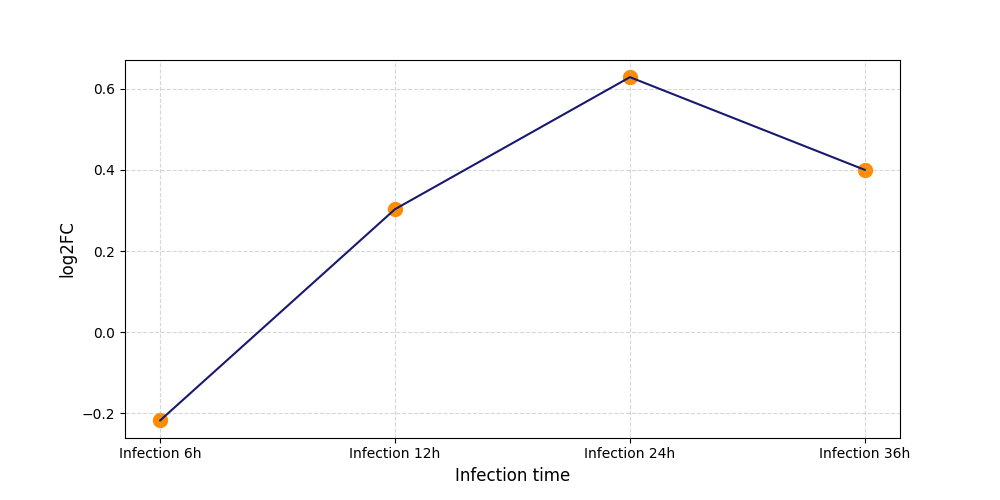
S1222
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
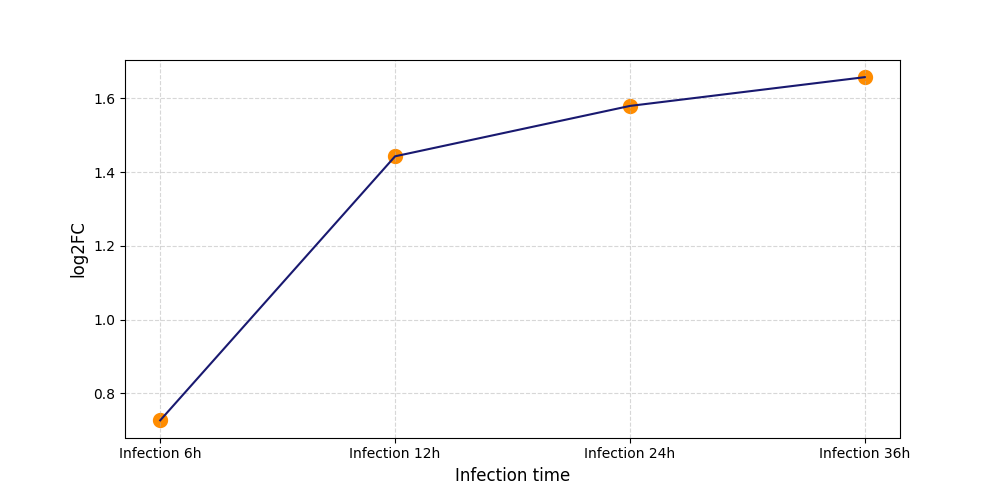
S1224
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
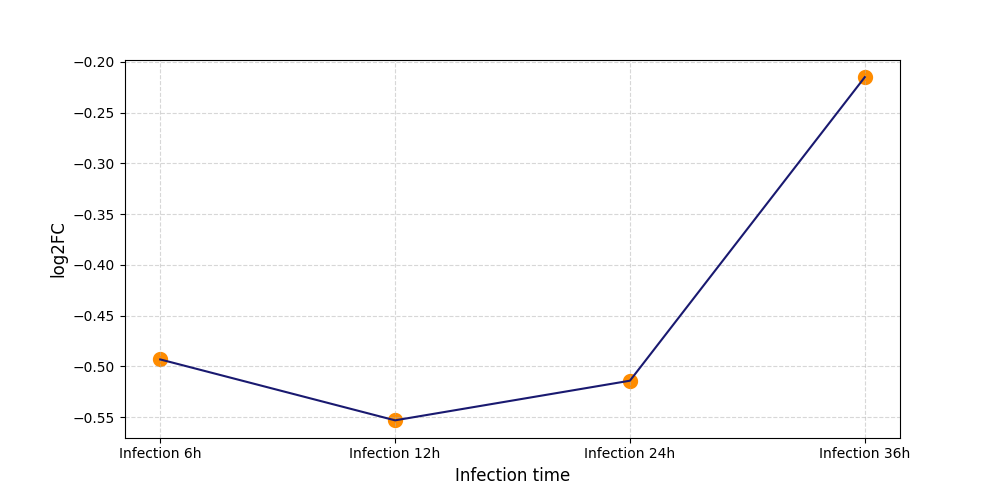
S1507
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
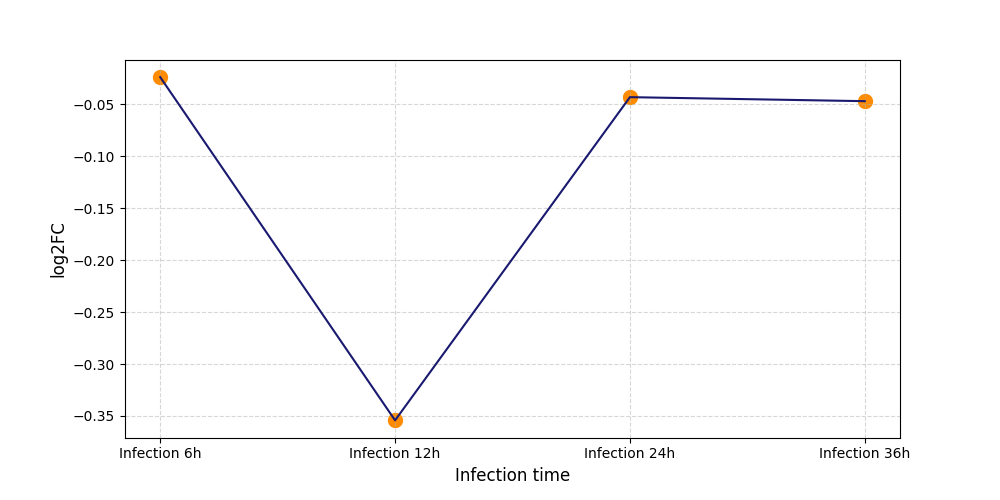
S2020
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
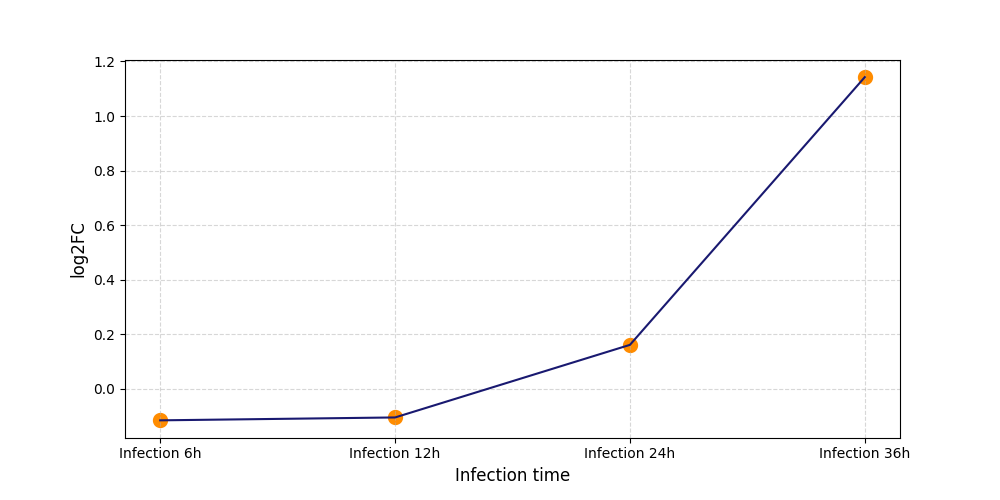
S411
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
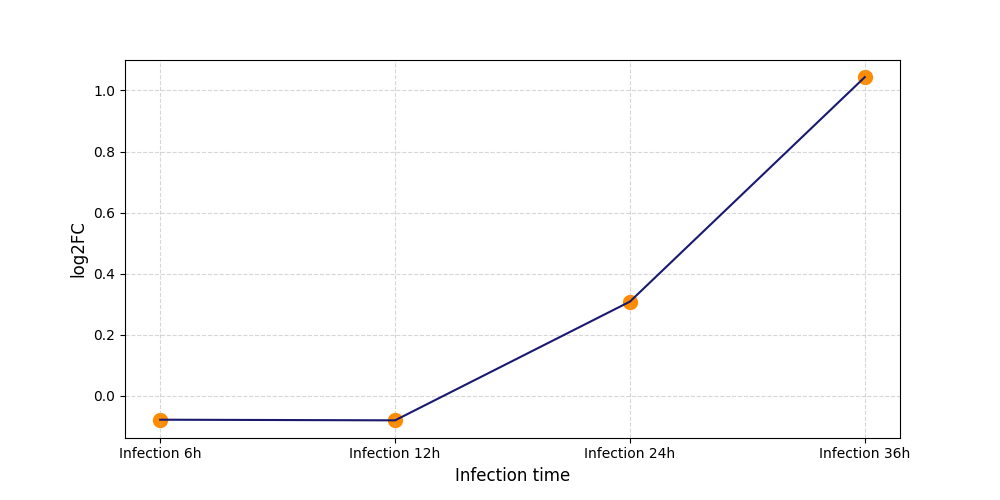
S419
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
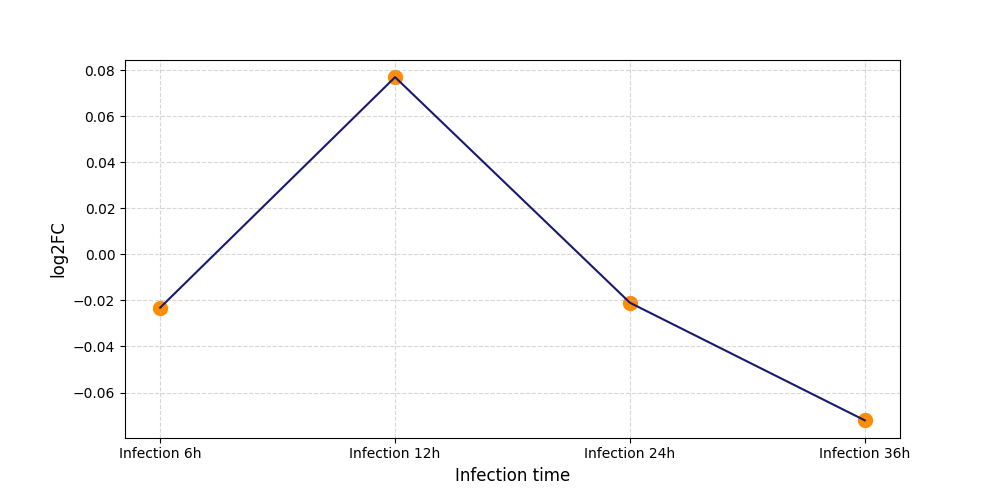
S598
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
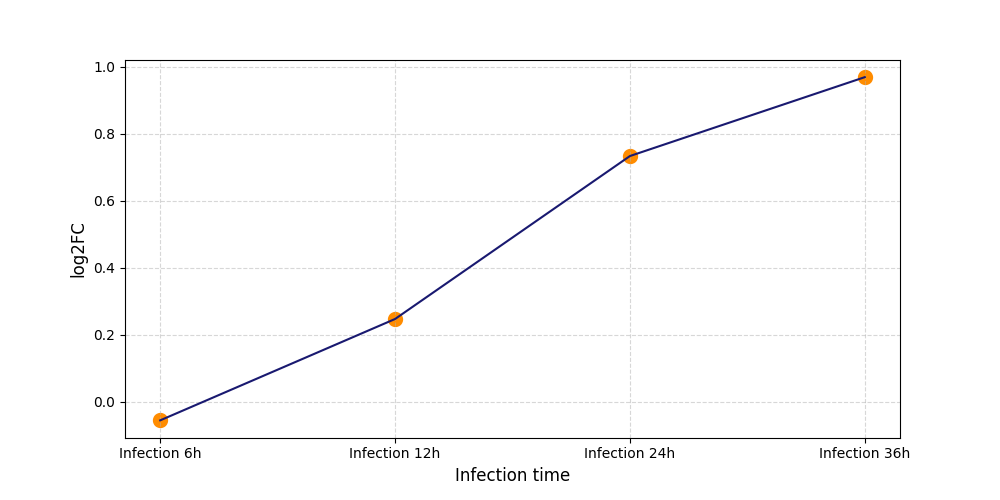
S6
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
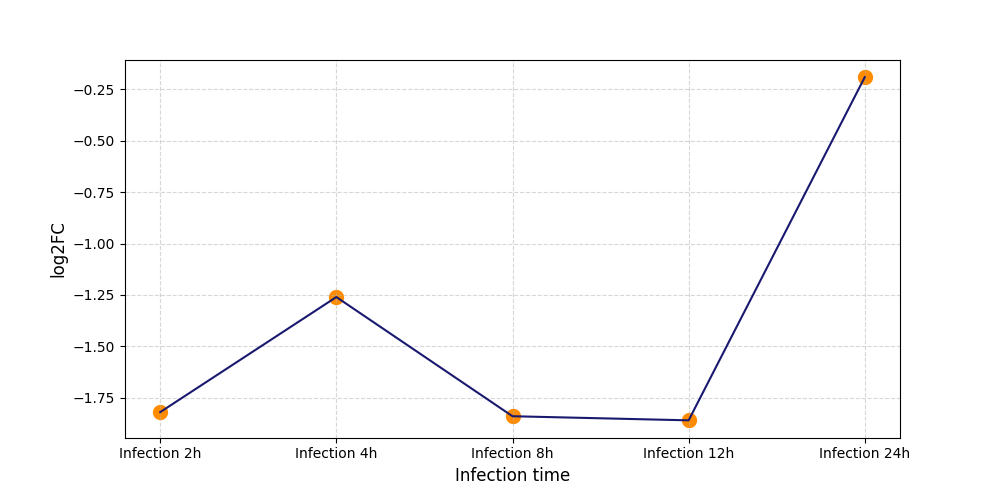
S666
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
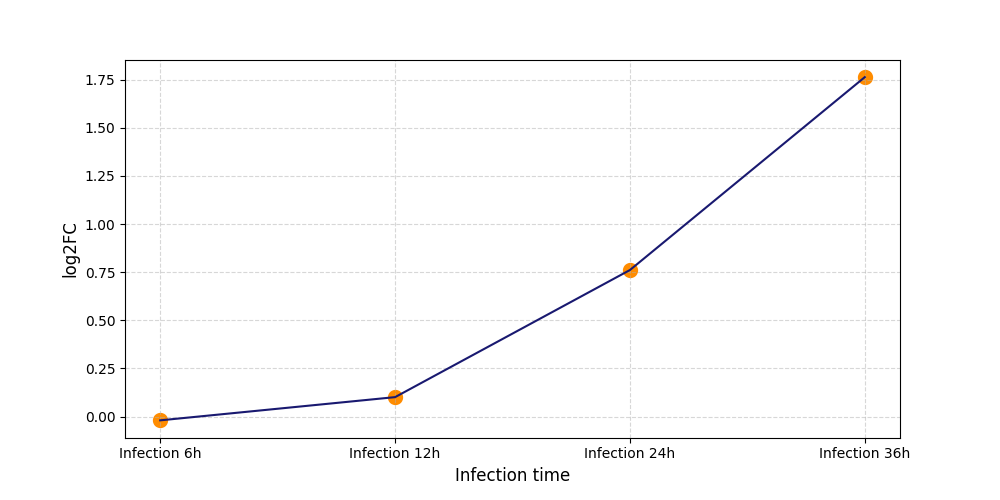
S666
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
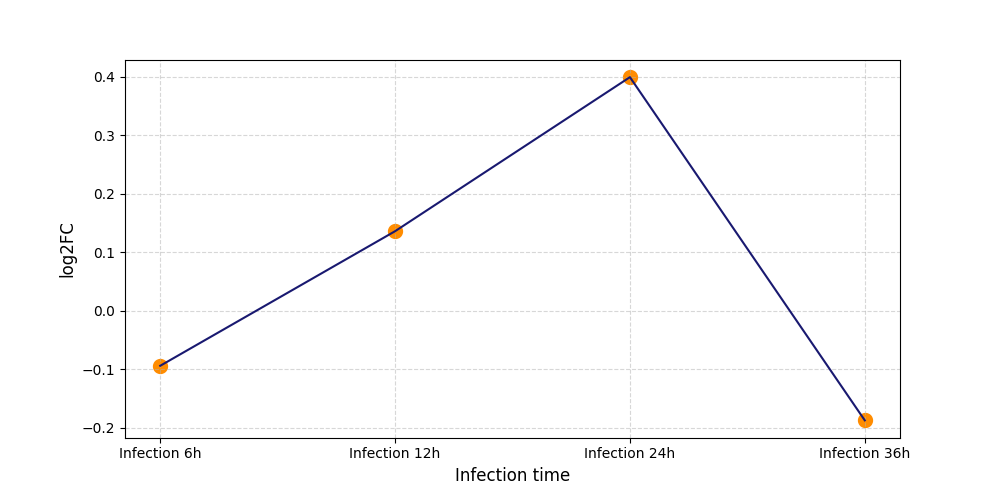
T413
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MASAVSPANLPAVLLQPRWKRVVGWSGPVPRPRHGHRAVAIKELIVVFGGGNEGIVDELHVYNTATNQWFIPAVRGDIPPGCAAYGFVCDGTRLLVFGGMVEYGKYSNDLYELQASRWEWKRLKAKTPKNGPPPCPRLGHSFSLVGNKCYLFGGLANDSEDPKNNIPRYLNDLYILELRPGSGVVAWDIPITYGVLPPPRESHTAVVYTEKDNKKSKLVIYGGMSGCRLGDLWTLDIDTLTWNKPSLSGVAPLPRSLHSATTIGNKMYVFGGWVPLVMDDVKVATHEKEWKCTNTLACLNLDTMAWETILMDTLEDNIPRARAGHCAVAINTRLYIWSGRDGYRKAWNNQVCCKDLWYLETEKPPPPARVQLVRANTNSLEVSWGAVATADSYLLQLQKYDIPATAATATSPTPNPVPSVPANPPKSPAPAAAAPAVQPLTQVGITLLPQAAPAPPTTTTIQVLPTVPGSSISVPTAARTQGVPAVLKVTGPQATTGTPLVTMRPASQAGKAPVTVTSLPAGVRMVVPTQSAQGTVIGSSPQMSGMAALAAAAAATQKIPPSSAPTVLSVPAGTTIVKTMAVTPGTTTLPATVKVASSPVMVSNPATRMLKTAAAQVGTSVSSATNTSTRPIITVHKSGTVTVAQQAQVVTTVVGGVTKTITLVKSPISVPGGSALISNLGKVMSVVQTKPVQTSAVTGQASTGPVTQIIQTKGPLPAGTILKLVTSADGKPTTIITTTQASGAGTKPTILGISSVSPSTTKPGTTTIIKTIPMSAIITQAGATGVTSSPGIKSPITIITTKVMTSGTGAPAKIITAVPKIATGHGQQGVTQVVLKGAPGQPGTILRTVPMGGVRLVTPVTVSAVKPAVTTLVVKGTTGVTTLGTVTGTVSTSLAGAGGHSTSASLATPITTLGTIATLSSQVINPTAITVSAAQTTLTAAGGLTTPTITMQPVSQPTQVTLITAPSGVEAQPVHDLPVSILASPTTEQPTATVTIADSGQGDVQPGTVTLVCSNPPCETHETGTTNTATTTVVANLGGHPQPTQVQFVCDRQEAAASLVTSTVGQQNGSVVRVCSNPPCETHETGTTNTATTATSNMAGQHGCSNPPCETHETGTTNTATTAMSSVGANHQRDARRACAAGTPAVIRISVATGALEAAQGSKSQCQTRQTSATSTTMTVMATGAPCSAGPLLGPSMAREPGGRSPAFVQLAPLSSKVRLSSPSIKDLPAGRHSHAVSTAAMTRSSVGAGEPRMAPVCESLQGGSPSTTVTVTALEALLCPSATVTQVCSNPPCETHETGTTNTATTSNAGSAQRVCSNPPCETHETGTTHTATTATSNGGTGQPEGGQQPPAGRPCETHQTTSTGTTMSVSVGALLPDATSSHRTVESGLEVAAAPSVTPQAGTALLAPFPTQRVCSNPPCETHETGTTHTATTVTSNMSSNQDPPPAASDQGEVESTQGDSVNITSSSAITTTVSSTLTRAVTTVTQSTPVPGPSVPPPEELQVSPGPRQQLPPRQLLQSASTALMGESAEVLSASQTPELPAAVDLSSTGEPSSGQESAGSAVVATVVVQPPPPTQSEVDQLSLPQELMAEAQAGTTTLMVTGLTPEELAVTAAAEAAAQAAATEEAQALAIQAVLQAAQQAVMGTGEPMDTSEAAATVTQAELGHLSAEGQEGQATTIPIVLTQQELAALVQQQQLQEAQAQQQHHHLPTEALAPADSLNDPAIESNCLNELAGTVPSTVALLPSTATESLAPSNTFVAPQPVVVASPAKLQAAATLTEVANGIESLGVKPDLPPPPSKAPMKKENQWFDVGVIKGTNVMVTHYFLPPDDAVPSDDDLGTVPDYNQLKKQELQPGTAYKFRVAGINACGRGPFSEISAFKTCLPGFPGAPCAIKISKSPDGAHLTWEPPSVTSGKIIEYSVYLAIQSSQAGGELKSSTPAQLAFMRVYCGPSPSCLVQSSSLSNAHIDYTTKPAIIFRIAARNEKGYGPATQVRWLQETSKDSSGTKPANKRPMSSPEMKSAPKKSKADGQ
Click to Show/Hide
|
|---|