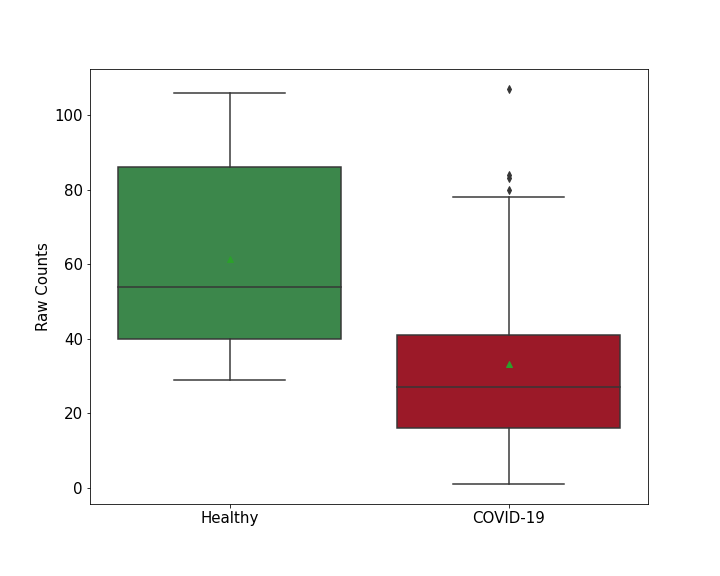
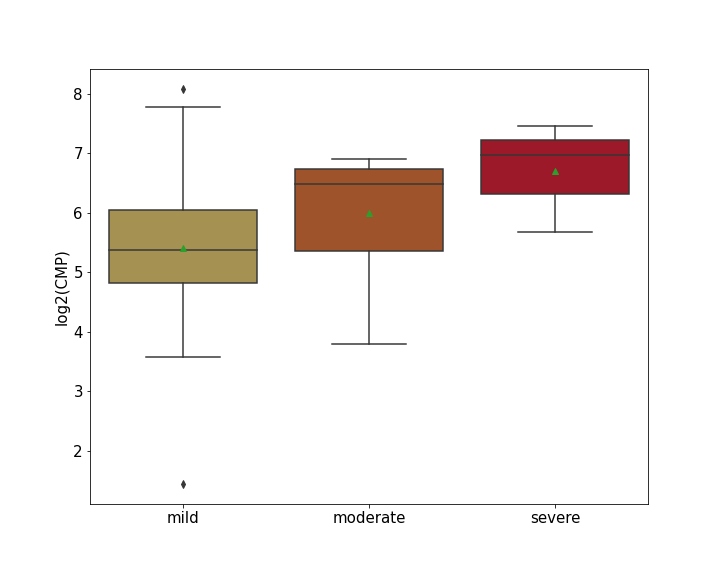
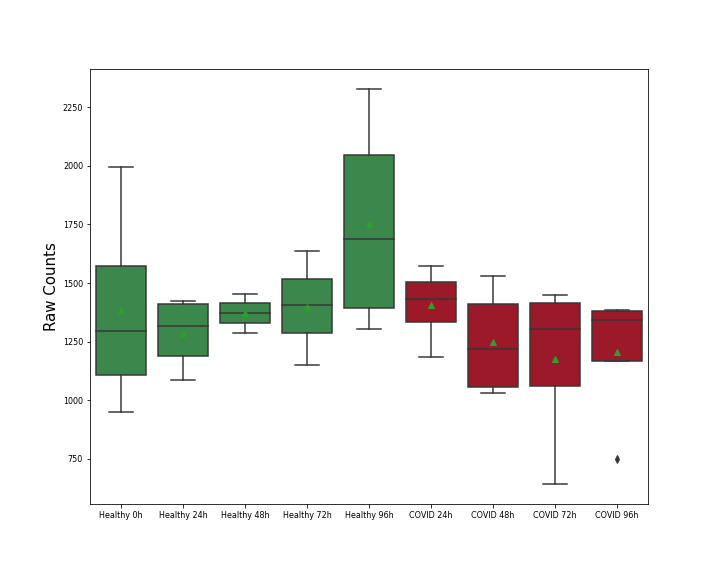
Details of Host Protein
| Host Protein General Information (ID: PT0615) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
L-lactate dehydrogenase B chain (LDHB)
|
Gene Name |
LDHB
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
LDHB_HUMAN
|
||||||
| Protein Families |
LDH/MDH superfamily
|
||||||||
| EC Number |
1.1.1.27
|
||||||||
| Subcellular Location |
Cytoplasm
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Interconverts simultaneously and stereospecifically pyruvateand lactate with concomitant interconversion of NADH and NAD (+).
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| HIF-1 signaling pathway | hsa04066 |
Pathway Map 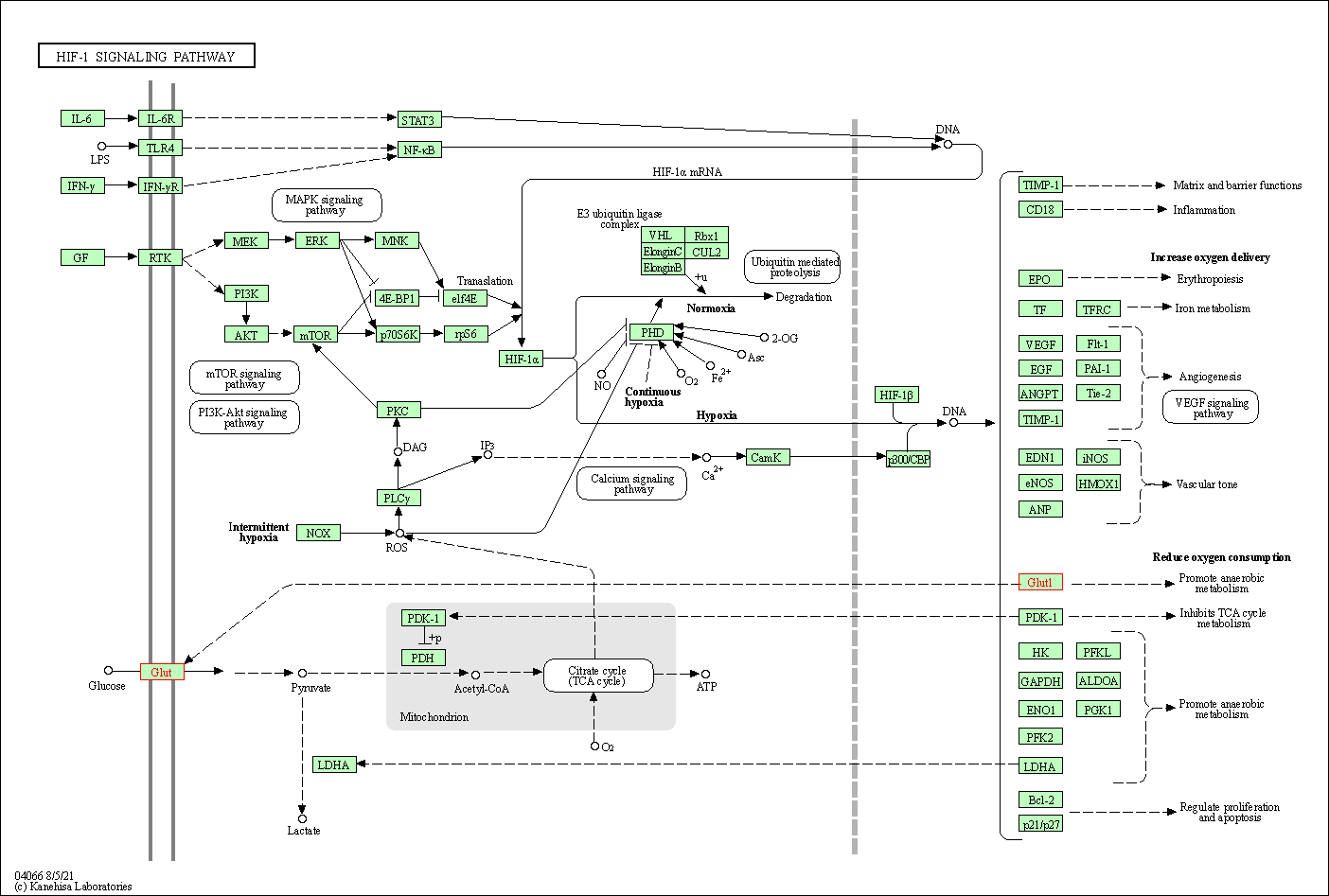
|
|||||||
| Cysteine and methionine metabolism | hsa00270 |
Pathway Map 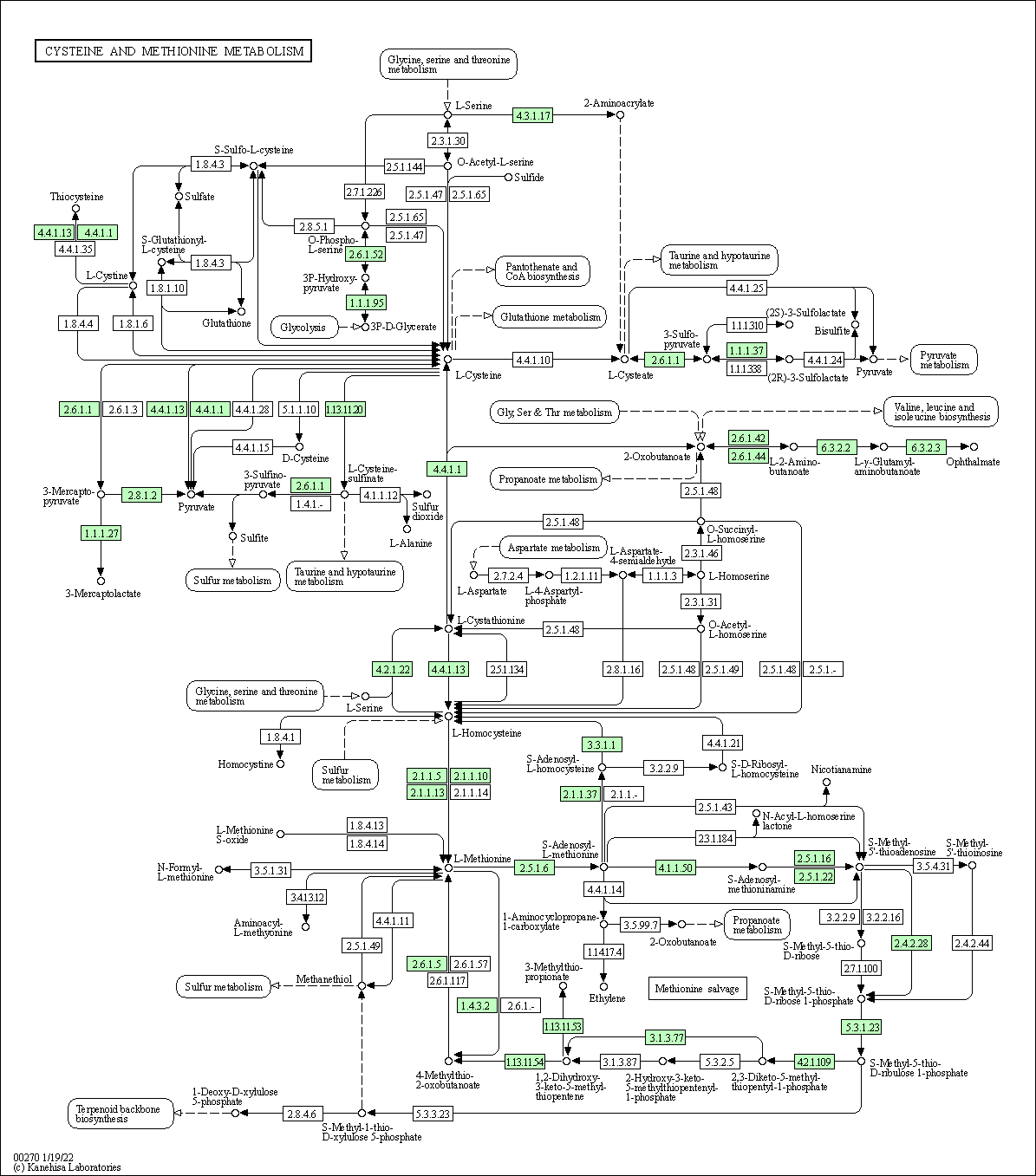
|
|||||||
| Central carbon metabolism in cancer | hsa05230 |
Pathway Map 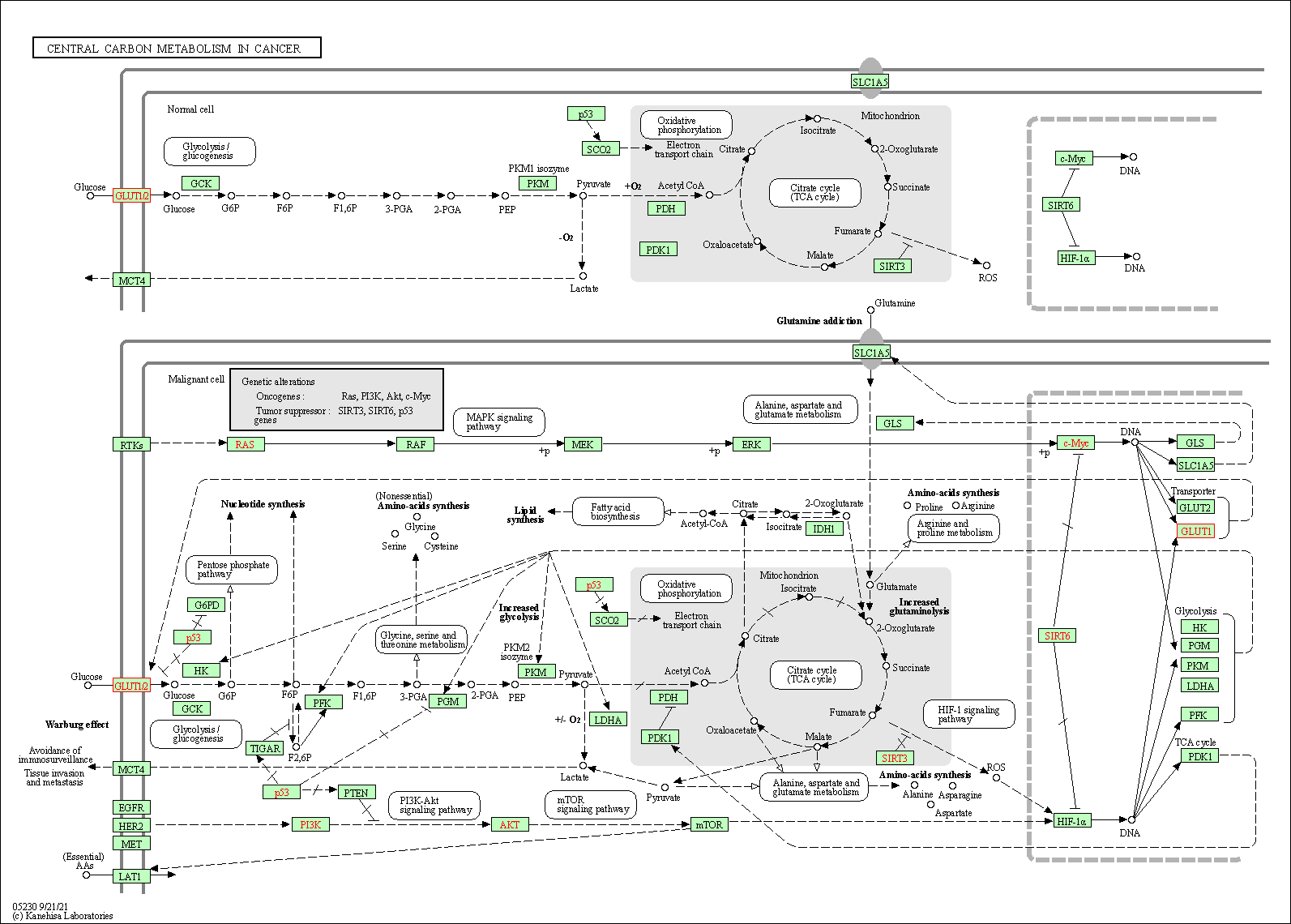
|
|||||||
| Glycolysis / Gluconeogenesis | hsa00010 |
Pathway Map 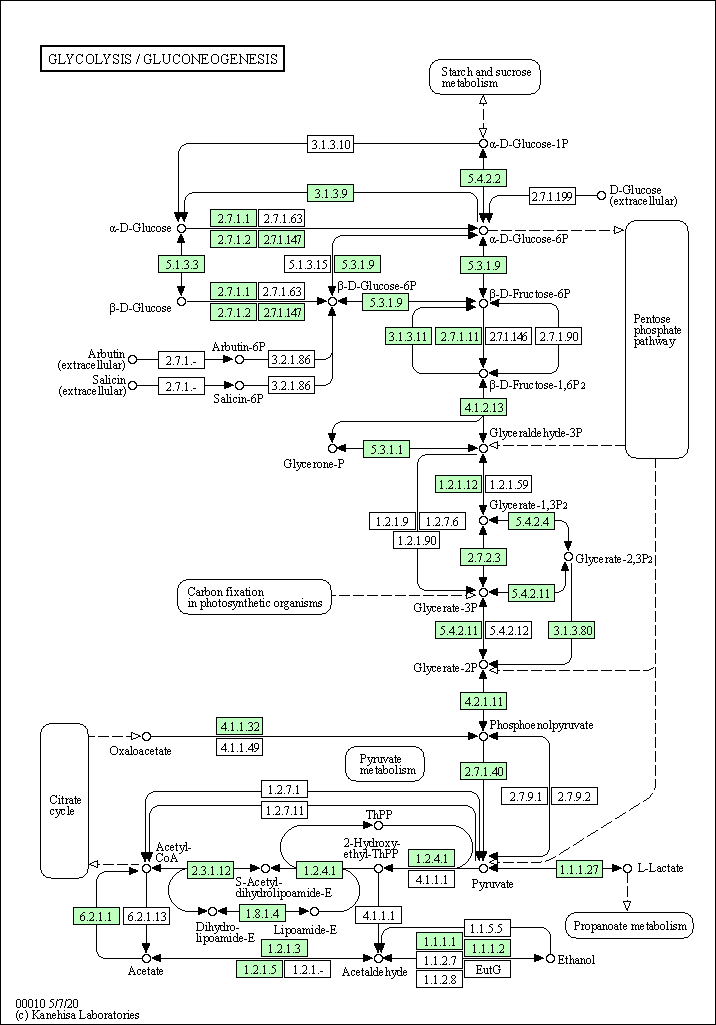
|
|||||||
| Pyruvate metabolism | hsa00620 |
Pathway Map 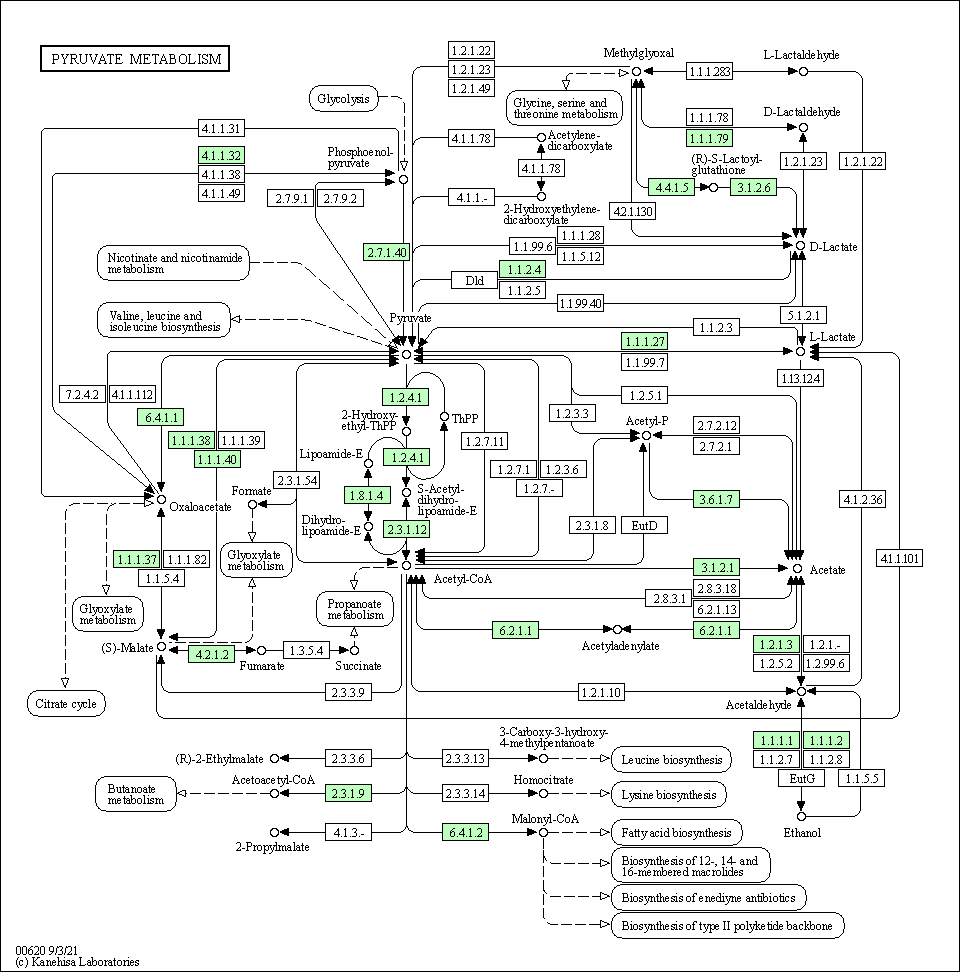
|
|||||||
| Propanoate metabolism | hsa00640 |
Pathway Map 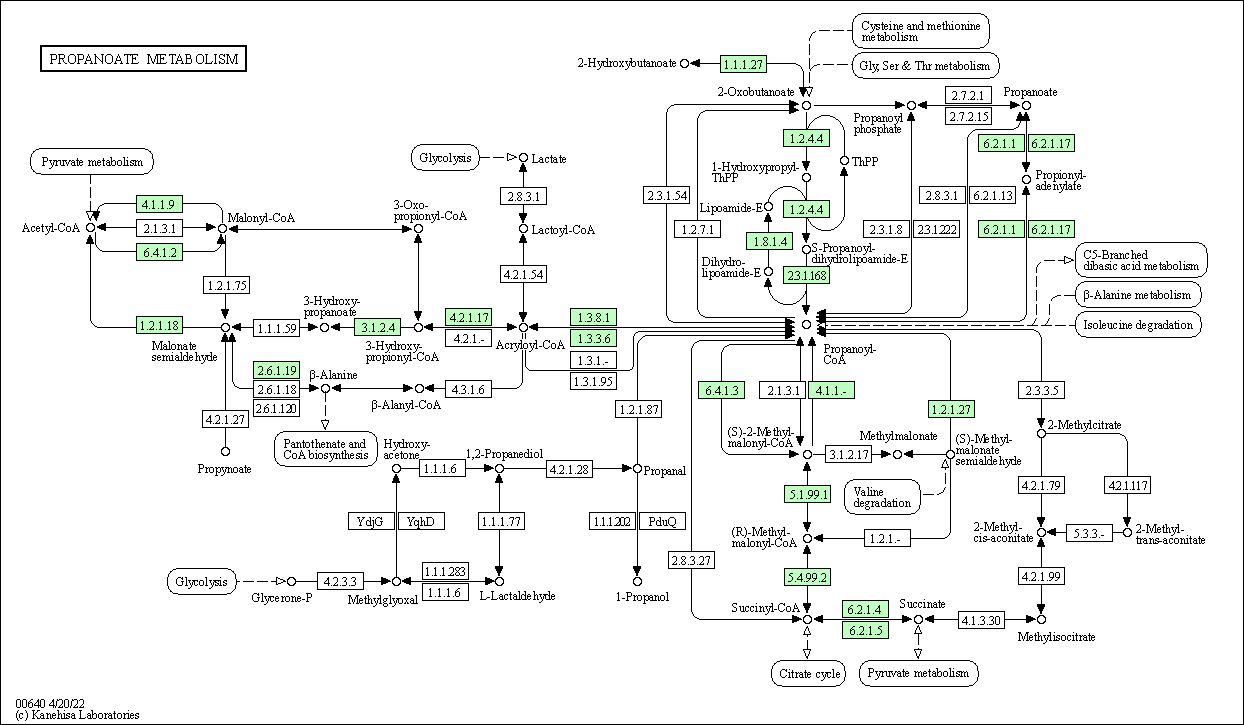
|
|||||||
| Glucagon signaling pathway | hsa04922 |
Pathway Map 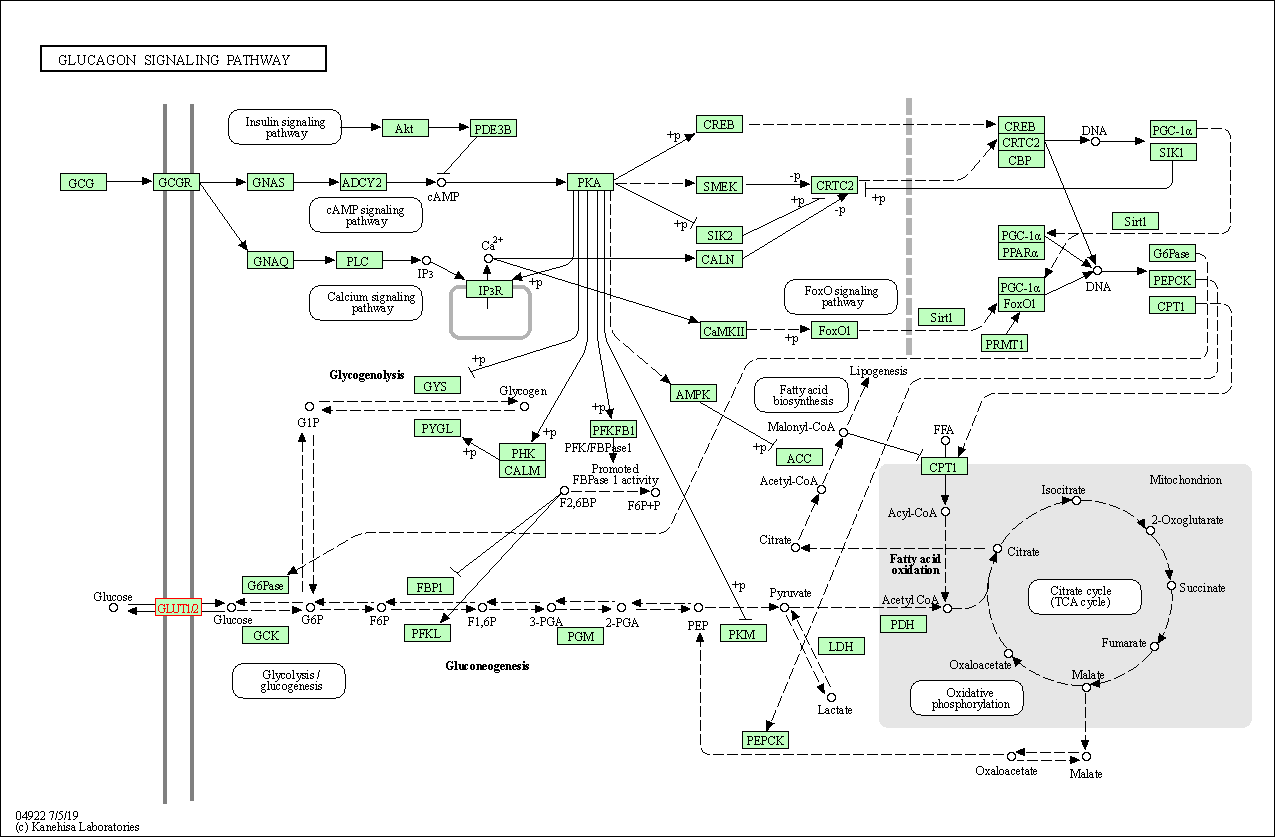
|
|||||||
| Metabolic pathways | hsa01100 |
Pathway Map 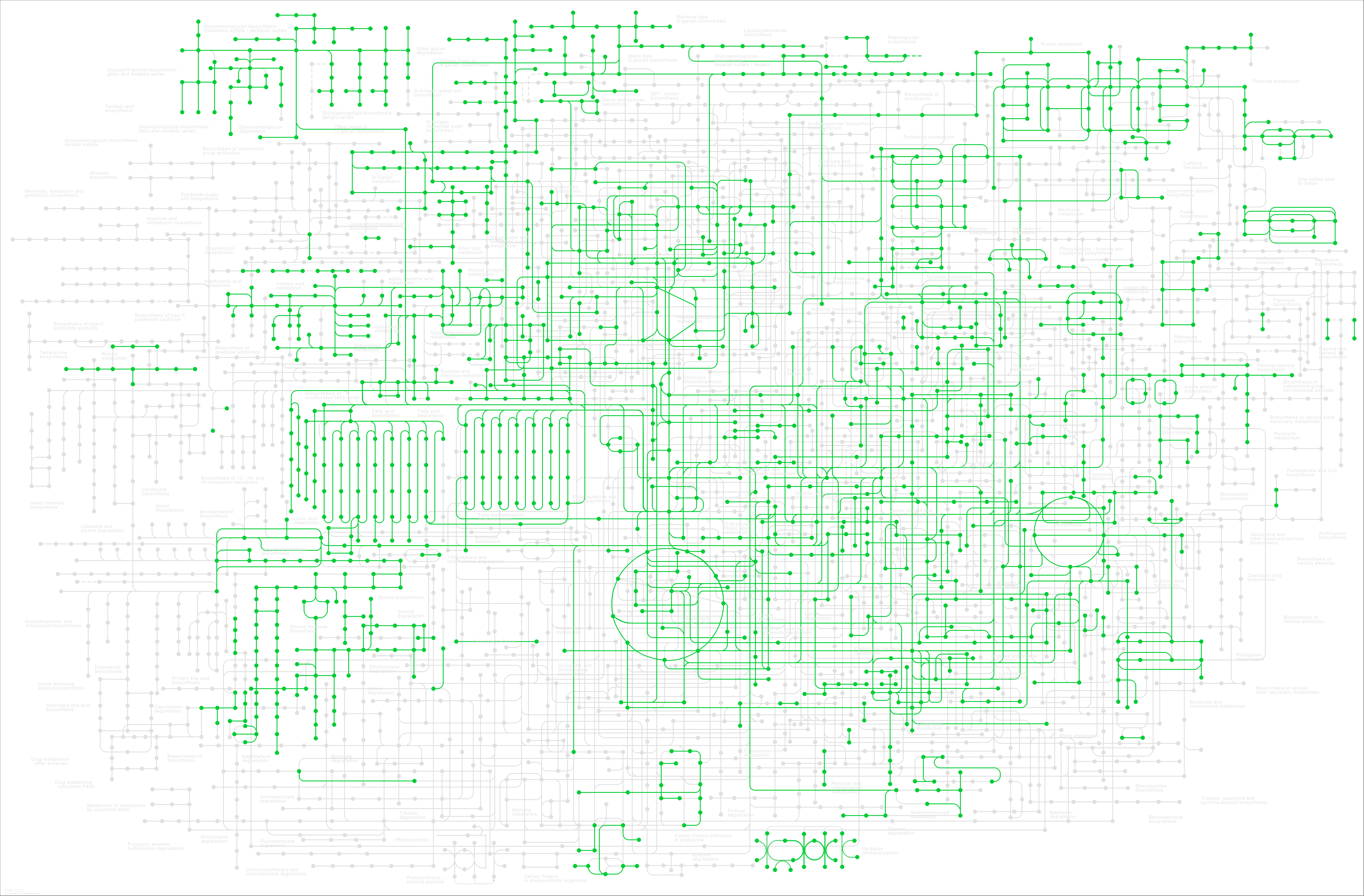
|
|||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Pro-viral | [1] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein LDHB in viral infection, LDHB protein knockout Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of LDHB leads to the decreased SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: ORF10 (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[2] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
ORF10
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Calu-3 cells (Human Lung Cancer Cell) (CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/France/IDF-220-95/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[3] | |||||||
| Strains Name |
hCoV-19/France/IDF-220-95/2020
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Direct interaction | ||||||||
| Infection Cells | HEK293 Cells (Human embryonic kidney cell) HEK293 Cells (Human embryonic kidney cell) (CVCL_0045 ) | ||||||||
| Cell Originated Tissue | Kidney | ||||||||
| Infection Time | 48 h | ||||||||
| Interaction Score | SAINT score ≥ 0.79 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
T18
[4] |
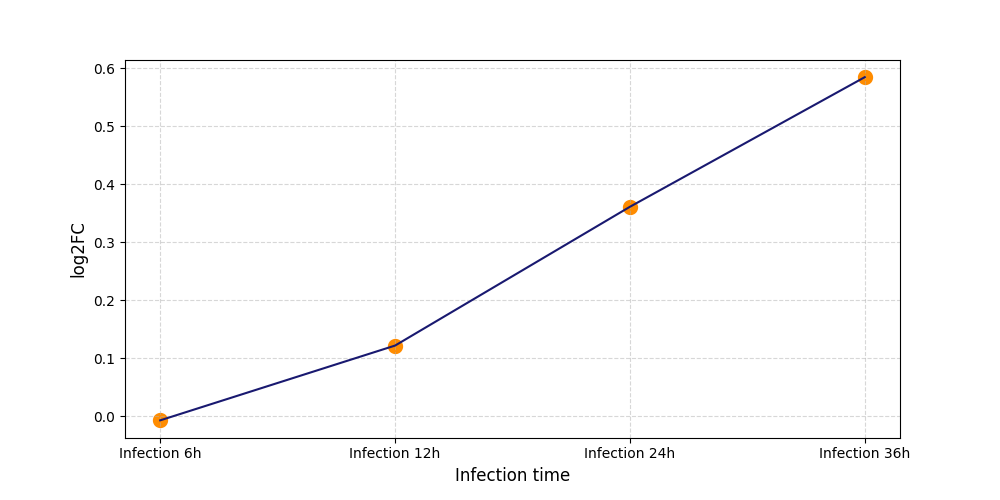 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MATLKEKLIAPVAEEEATVPNNKITVVGVGQVGMACAISILGKSLADELALVDVLEDKLKGEMMDLQHGSLFLQTPKIVADKDYSVTANSKIVVVTAGVRQQEGESRLNLVQRNVNVFKFIIPQIVKYSPDCIIIVVSNPVDILTYVTWKLSGLPKHRVIGSGCNLDSARFRYLMAEKLGIHPSSCHGWILGEHGDSSVAVWSGVNVAGVSLQELNPEMGTDNDSENWKEVHKMVVESAYEVIKLKGYTNWAIGLSVADLIESMLKNLSRIHPVSTMVKGMYGIENEVFLSLPCILNARGLTSVINQKLKDDEVAQLKKSADTLWDIQKDLKDL
Click to Show/Hide
|
|---|