Details of Host Protein
| Host Protein General Information (ID: PT0710) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
Paraspeckle component 1 (PSPC1)
|
Gene Name |
PSPC1
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
PSPC1_HUMAN
|
||||||
| Protein Families |
PSPC family
|
||||||||
| Subcellular Location |
Nucleus; nucleolus
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Regulates, cooperatively with NONO and SFPQ, androgenreceptor-mediated gene transcription activity in Sertoli cell line. Binds to poly (A), poly (G) and poly (U) RNA homopolymers. Regulates the circadian clock by repressing the transcriptionalactivator activity of the CLOCK-ARNTL/BMAL1 heterodimer. Together with NONO, required for the formation of nuclearparaspeckles. Plays a role in the regulation of DNA virus-mediatedinnate immune response by assembling into the HDP-RNP complex, acomplex that serves as a platform for IRF3 phosphorylation andsubsequent innate immune response activation through the cGAS-STINGpathway.
[1-2]
Click to Show/Hide
|
||||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Pro-viral | [3] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein PSPC1 in viral infection, PSPC1 protein knockout Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of PSPC1 leads to the decreased SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: 3'-UTR (hCoV-19/IPBCAMS-YL01/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[4] | |||||||
| Strains Name |
hCoV-19/IPBCAMS-YL01/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potiential to be direct binder | ||||||||
| Interaction Binding Type | Single stranded RNA binding | ||||||||
| Infection Cells | Huh7.5.1 cells (Hepatocyte derived cellular carcinoma cell) (CVCL_E049 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 30 h | ||||||||
| Interaction Score | MIST = 0.68551467 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||
| RNA Region: 5'-UTR (hCoV-19/South Korea/KCDC03/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/South Korea/KCDC03/2020
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Directly bind to SARS-CoV-23 RNA's 5' UTR region | ||||||||
| Infection Cells | Vero cells (epithelial kidney cell) Vero cells (epithelial kidney cell) (CVCL_0059 ) | ||||||||
| Cell Originated Tissue | kidney | ||||||||
| Infection Time | 24 h | ||||||||
| Interaction Score | P-adjust < 0.05 | ||||||||
| Method Description | Modified RNA antisense purification coupled with mass spectrometry (RAP-MS); label-free quantification (LFQ); liquid chromatography with tandem mass spectrometry (LC-MS/MS) | ||||||||
| RNA Region: Not Specified Virus Region (hCov-OC43/VR-759 Quebec/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCov-OC43/VR-759 Quebec/2019
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus OC43 (HCov-OC43)
|
||||||||
| Infection Cells | HCT-8 cells (Human ileocaecal adenocarcinoma cell) HCT-8 cells (Human ileocaecal adenocarcinoma cell) (CVCL_2478 ) | ||||||||
| Cell Originated Tissue | Ileocecum | ||||||||
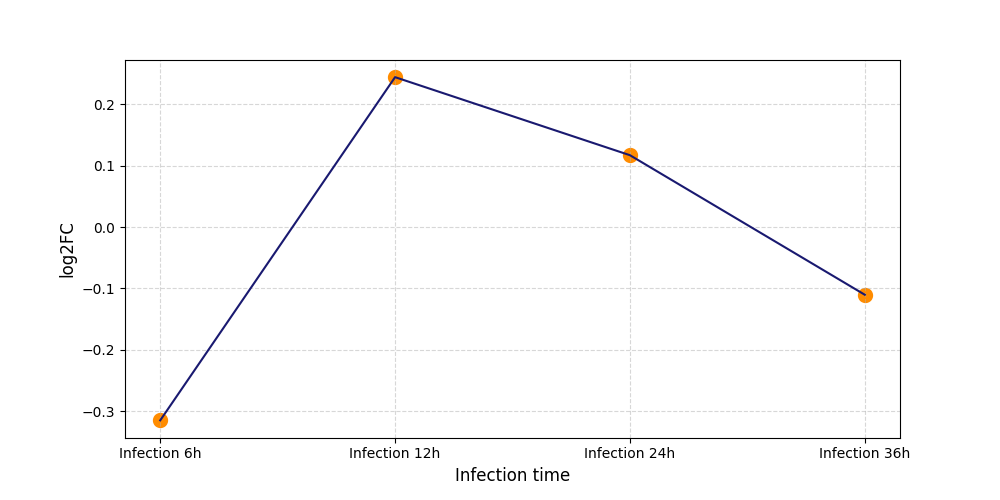
| Infection Time | 12h; 24; 36h; 48h | ||||||||
| Interaction Score | p_value < 0.05; FDR < 10% | ||||||||
| Method Description | Modified RNA antisense purification coupled with mass spectrometry (RAP-MS); label-free quantification (LFQ); liquid chromatography with tandem mass spectrometry (LC-MS/MS) | ||||||||
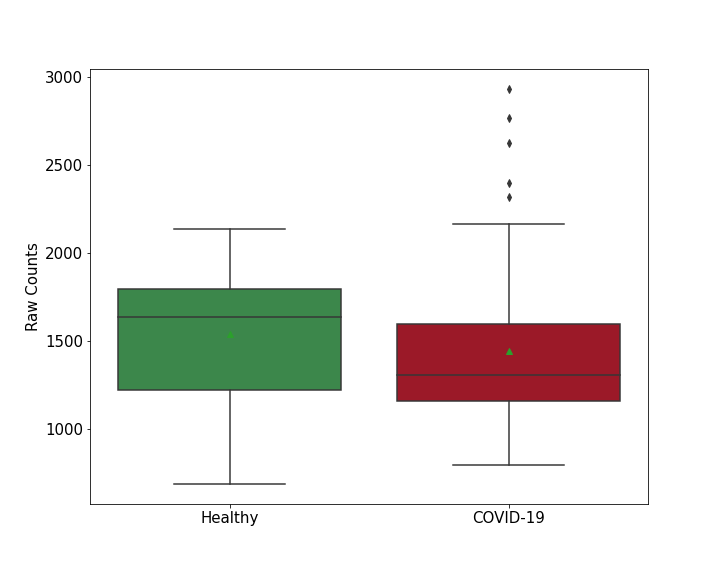
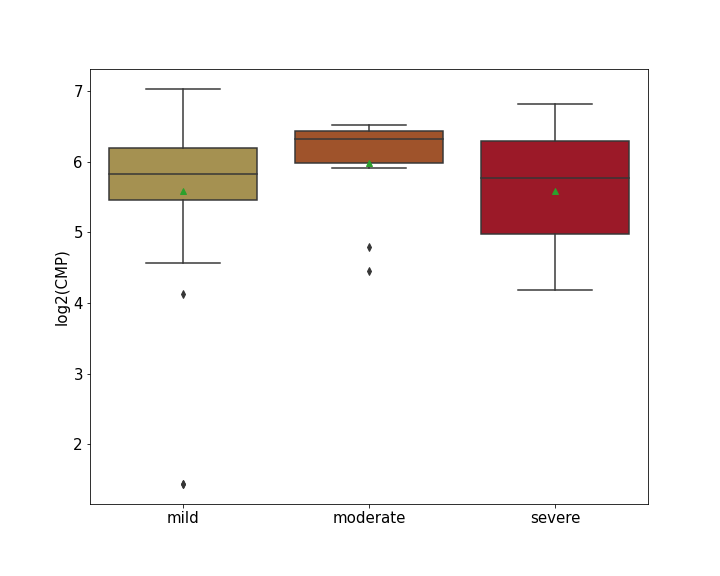
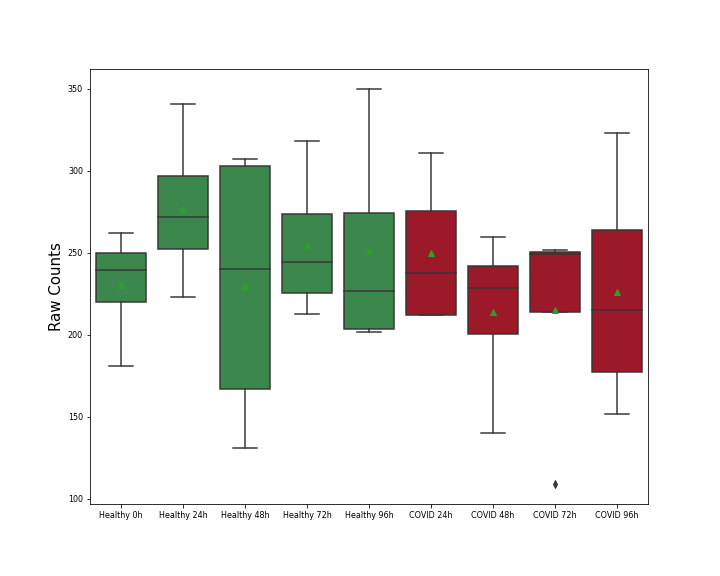
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
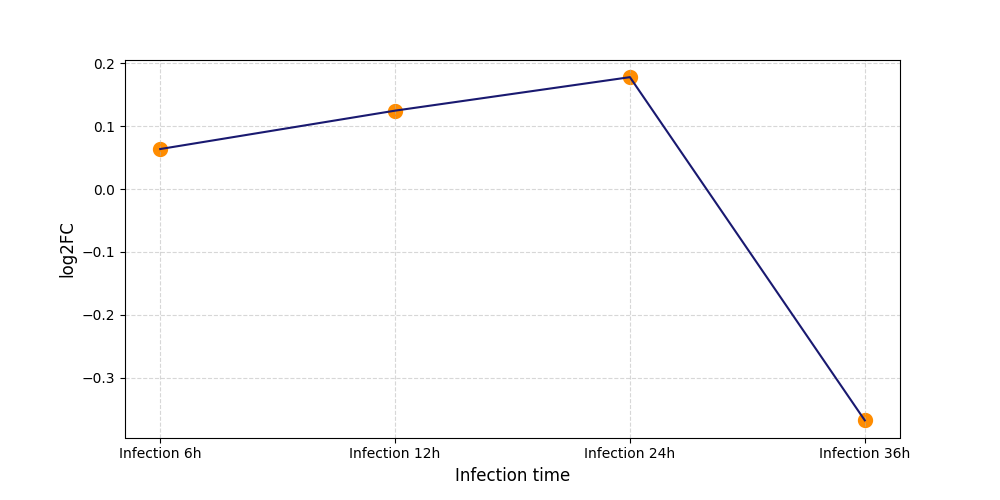
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S473
[6] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
S477
[6] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MMLRGNLKQVRIEKNPARLRALESAVGESEPAAAAAMALALAGEPAPPAPAPPEDHPDEEMGFTIDIKSFLKPGEKTYTQRCRLFVGNLPTDITEEDFKRLFERYGEPSEVFINRDRGFGFIRLESRTLAEIAKAELDGTILKSRPLRIRFATHGAALTVKNLSPVVSNELLEQAFSQFGPVEKAVVVVDDRGRATGKGFVEFAAKPPARKALERCGDGAFLLTTTPRPVIVEPMEQFDDEDGLPEKLMQKTQQYHKEREQPPRFAQPGTFEFEYASRWKALDEMEKQQREQVDRNIREAKEKLEAEMEAARHEHQLMLMRQDLMRRQEELRRLEELRNQELQKRKQIQLRHEEEHRRREEEMIRHREQEELRRQQEGFKPNYMENREQEMRMGDMGPRGAINMGDAFSPAPAGNQGPPPMMGMNMNNRATIPGPPMGPGPAMGPEGAANMGTPMMPDNGAVHNDRFPQGPPSQMGSPMGSRTGSETPQAPMSGVGPVSGGPGGFGRGSQGGNFEGPNKRRRY
Click to Show/Hide
|
|---|