Details of Host Protein
| Host Protein General Information (ID: PT0998) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
Histone-glutamine methyltransferase (FBL)
|
Gene Name |
FBL
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
FBRL_HUMAN
|
||||||
| Protein Families |
Methyltransferase superfamily
|
||||||||
| EC Number |
2.1.1.-
|
||||||||
| Subcellular Location |
Nucleus; nucleolus Nucleus; nucleoplasm
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
S-adenosyl-L-methionine-dependent methyltransferase that hasthe ability to methylate both RNAs and proteins. Involved in pre-rRNA processing bycatalyzing the site-specific 2'-hydroxyl methylation of ribose moietiesin pre-ribosomal RNA. Site specificity is provided bya guide RNA that base pairs with the substrate. Methylation occurs at a characteristic distance from the sequenceinvolved in base pairing with the guide RNA. Probablycatalyzes 2'-O-methylation of U6 snRNAs in box C/D RNP complexes. U6 snRNA 2'-O-methylation is required for mRNAsplicing fidelity. Also acts as a proteinmethyltransferase by mediating methylation of 'Gln-105' of histone H2A (H2AQ104me), a modification that impairs binding of the FACT complexand is specifically present at 35S ribosomal DNA locus.
[1-2]
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| Ribosome biogenesis in eukaryotes | hsa03008 |
Pathway Map 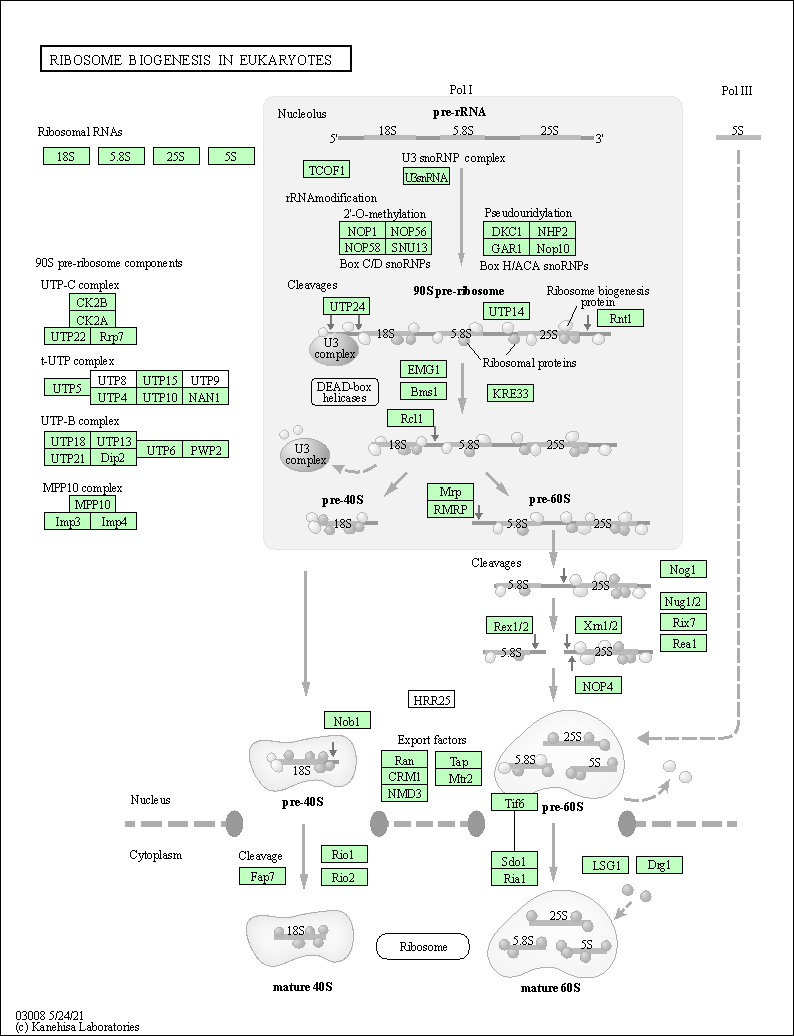
|
|||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Anti-viral | [3] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
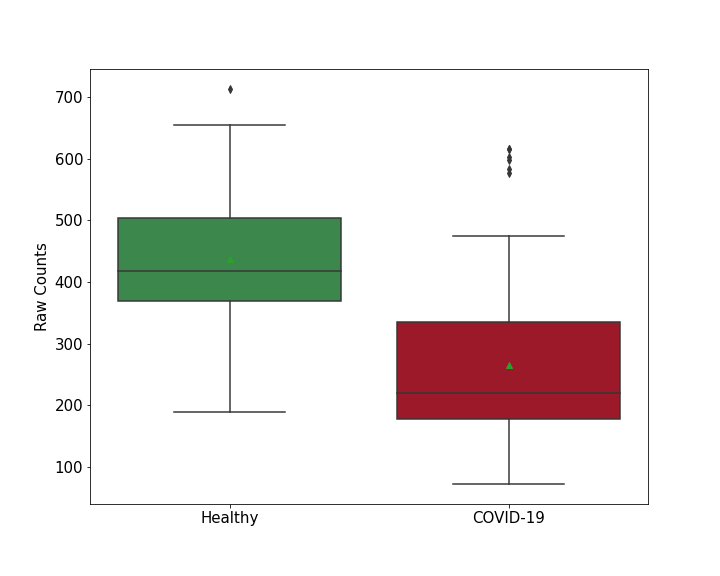
| Method Description | To detect the role of host protein FBL in viral infection, FBL protein knockdown Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of FBL increases SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: 5'-UTR (hCoV-19/IPBCAMS-YL01/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[4] | |||||||
| Strains Name |
hCoV-19/IPBCAMS-YL01/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7.5.1 cells (Hepatocyte derived cellular carcinoma cell) (CVCL_E049 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 30 h | ||||||||
| Interaction Score | Affinity = 0.941 | ||||||||
| Method Description | PrismNet; pull-down western (STAR methods); Mass spectrometry | ||||||||
| RNA Region: ORF10 (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF10
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) (CVCL_0336;CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Liver; Lung | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
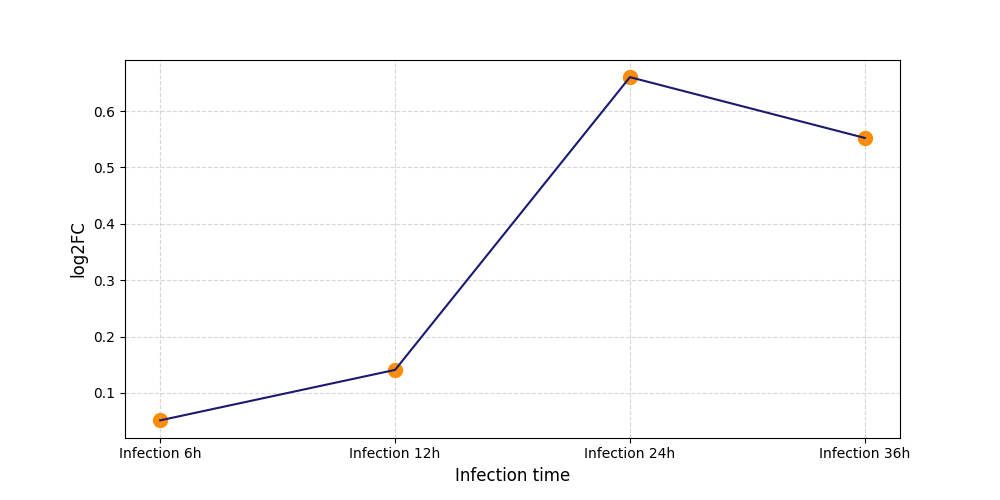
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S124
[6] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MKPGFSPRGGGFGGRGGFGDRGGRGGRGGFGGGRGRGGGFRGRGRGGGGGGGGGGGGGRGGGGFHSGGNRGRGRGGKRGNQSGKNVMVEPHRHEGVFICRGKEDALVTKNLVPGESVYGEKRVSISEGDDKIEYRAWNPFRSKLAAAILGGVDQIHIKPGAKVLYLGAASGTTVSHVSDIVGPDGLVYAVEFSHRSGRDLINLAKKRTNIIPVIEDARHPHKYRMLIAMVDVIFADVAQPDQTRIVALNAHTFLRNGGHFVISIKANCIDSTASAEAVFASEVKKMQQENMKPQEQLTLEPYERDHAVVVGVYRPPPKVKN
Click to Show/Hide
|
|---|