Details of Host Protein
| Host Protein General Information (ID: PT0220) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
Suppressor of var1 3-like protein 1 (SUV3)
|
Gene Name |
SUPV3L1
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
SUV3_HUMAN
|
||||||
| Protein Families |
Helicase family
|
||||||||
| EC Number |
3.6.4.13
|
||||||||
| Subcellular Location |
Mitochondrion; Nucleus
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Major helicase player in mitochondrial RNA metabolism. Component of the mitochondrial degradosome (mtEXO) complex, thatdegrades 3' overhang double-stranded RNA with a 3'-to-5' directionalityin an ATP-dependent manner. Involved in the degradation of non-codingmitochondrial transcripts (MT-ncRNA) and tRNA-like molecules. ATPase and ATP-dependent multisubstrate helicase, able to unwind double-stranded (ds) DNA and RNA, and RNA/DNAheteroduplexes in the 5'-to-3' direction. Plays a role in the RNAsurveillance system in mitochondria; regulates the stability of maturemRNAs, the removal of aberrantly formed mRNAs and the rapid degradationof non coding processing intermediates. Also implicated inrecombination and chromatin maintenance pathways. May protect cellsfrom apoptosis. Associates with mitochondrial DNA.
[1-8]
Click to Show/Hide
|
||||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Anti-viral | [9] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein SUPV3L1 in viral infection, SUPV3L1 protein knockout Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
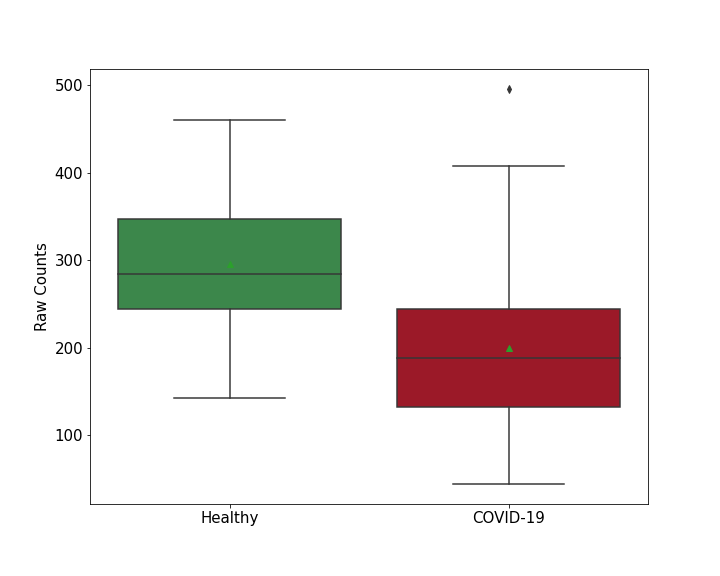
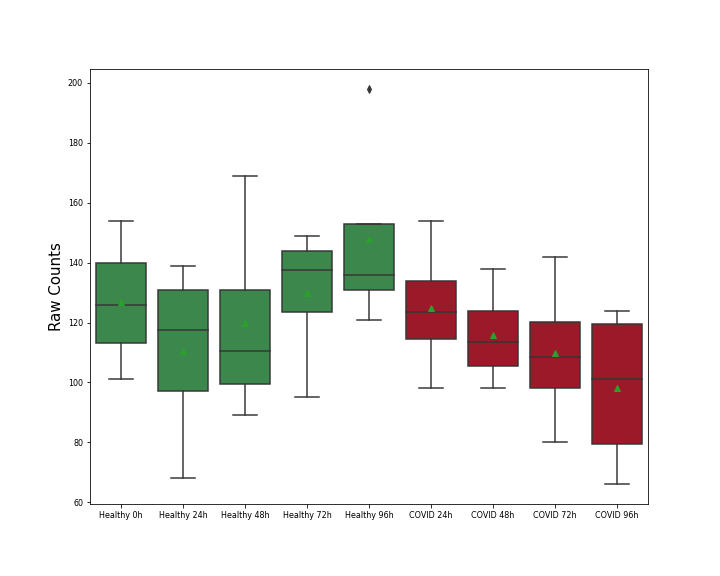
| Results | It is reported that knockout of SUPV3L1 increases SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: E region (hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Wuhan/HBCDC-HB-04/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Wuhan/HBCDC-HB-04/2020
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Philippines/PH-PGC-104159/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Philippines/PH-PGC-104159/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Myanmar/DSMRC-065/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Myanmar/DSMRC-065/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Malaysia/MGI_GS0622/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Malaysia/MGI_GS0622/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Ethiopia/ILRI_COVM02835/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Ethiopia/ILRI_COVM02835/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/ElSalvador/INC-LNSP-153/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/ElSalvador/INC-LNSP-153/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Chile/AT-59206/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Chile/AT-59206/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Canada/UN-276648/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Canada/UN-276648/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Bonaire/BQ-RIVM-101067/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Bonaire/BQ-RIVM-101067/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Bangladesh/CHRF-GSB17-117/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Bangladesh/CHRF-GSB17-117/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Australia/SA60463/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Australia/SA60463/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Wuhan/HBCDC-HB-04/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Wuhan/HBCDC-HB-04/2020
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Philippines/PH-PGC-104159/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Philippines/PH-PGC-104159/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Myanmar/DSMRC-065/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Myanmar/DSMRC-065/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Malaysia/MGI_GS0622/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Malaysia/MGI_GS0622/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Ethiopia/ILRI_COVM02835/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Ethiopia/ILRI_COVM02835/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/ElSalvador/INC-LNSP-153/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/ElSalvador/INC-LNSP-153/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Chile/AT-59206/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Chile/AT-59206/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Canada/UN-276648/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Canada/UN-276648/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Bonaire/BQ-RIVM-101067/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Bonaire/BQ-RIVM-101067/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Bangladesh/CHRF-GSB17-117/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Bangladesh/CHRF-GSB17-117/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Australia/SA60463/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Australia/SA60463/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Scotland/QEUH-13A999F/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Scotland/QEUH-13A999F/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Scotland/EDB14267/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/Scotland/EDB14267/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/QEUH-1269F5E/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/England/QEUH-1269F5E/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/MILK-F88D46/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/England/MILK-F88D46/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/MILK-F88C2B/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/England/MILK-F88C2B/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/MILK-1205929/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/England/MILK-1205929/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/MILK-11F02FA/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/England/MILK-11F02FA/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/LOND-12F444A/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/England/LOND-12F444A/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/CAMC-13B6BB7/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/England/CAMC-13B6BB7/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/CAMC-12DF946/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/England/CAMC-12DF946/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-SARS/Manitoba/2003 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-SARS/Manitoba/2003
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome-related coronavirus (HCoV-SARS)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-OC43/VR-759 Quebec/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCov-OC43/VR-759 Quebec/2019
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus OC43 (HCov-OC43)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-NL63/Amsterdam/2004 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCov-NL63/Amsterdam/2004
|
||||||||
| Strains Family |
Alpha
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human Coronavirus NL63 (HCov-NL63)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-MERS/Jeddah/2012 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCov-MERS/Jeddah/2012
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Middle East respiratory syndrome-related coronavirus (HCov-MERS)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-HKU1/Hong Kong/2004 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-HKU1/Hong Kong/2004
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus HKU1 (HCoV-HKU1)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-229E/Würzburg/2000 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCov-229E/Würzburg/2000
|
||||||||
| Strains Family |
Alpha
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus 229E (HCov-229E)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||