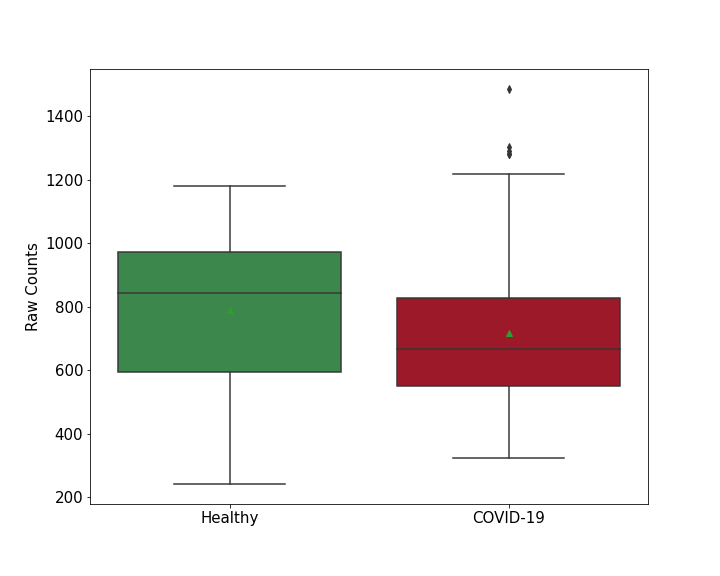
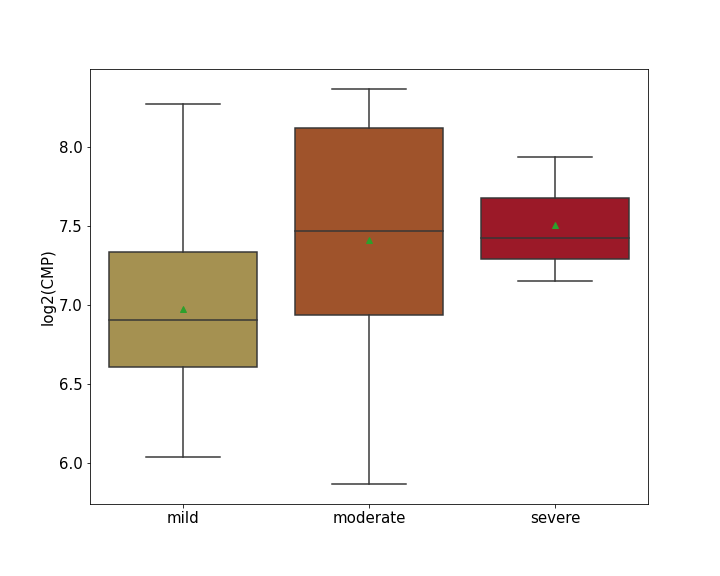
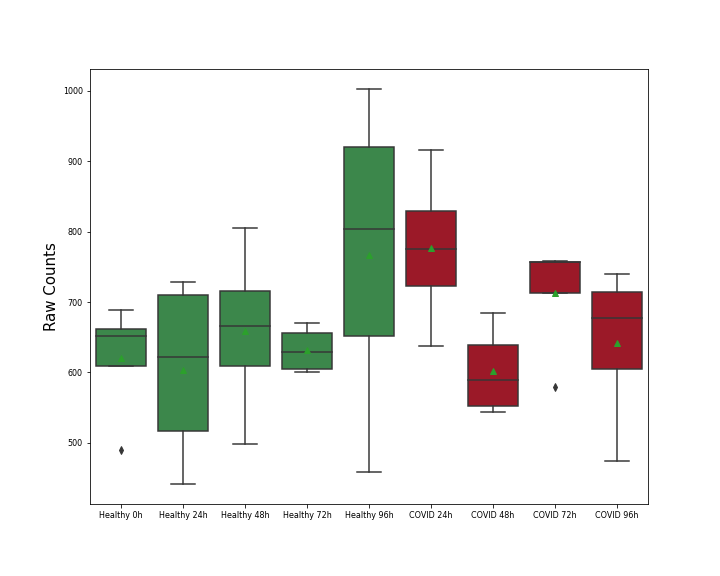
Details of Host Protein
| Host Protein General Information (ID: PT0476) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
D-fructose-6-phosphate amidotransferase 1 (GFPT1)
|
Gene Name |
GFPT1
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
GFPT1_HUMAN
|
||||||
| EC Number |
2.6.1.16
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Controls the flux of glucose into the hexosamine pathway. Most likely involved in regulating the availability of precursors forN- and O-linked glycosylation of proteins. Regulates the circadianexpression of clock genes ARNTL/BMAL1 and CRY1. Has arole in fine tuning the metabolic fluctuations of cytosolic UDP-GlcNAcand its effects on hyaluronan synthesis that occur during tissueremodeling.
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| Alanine, aspartate and glutamate metabolism | hsa00250 |
Pathway Map 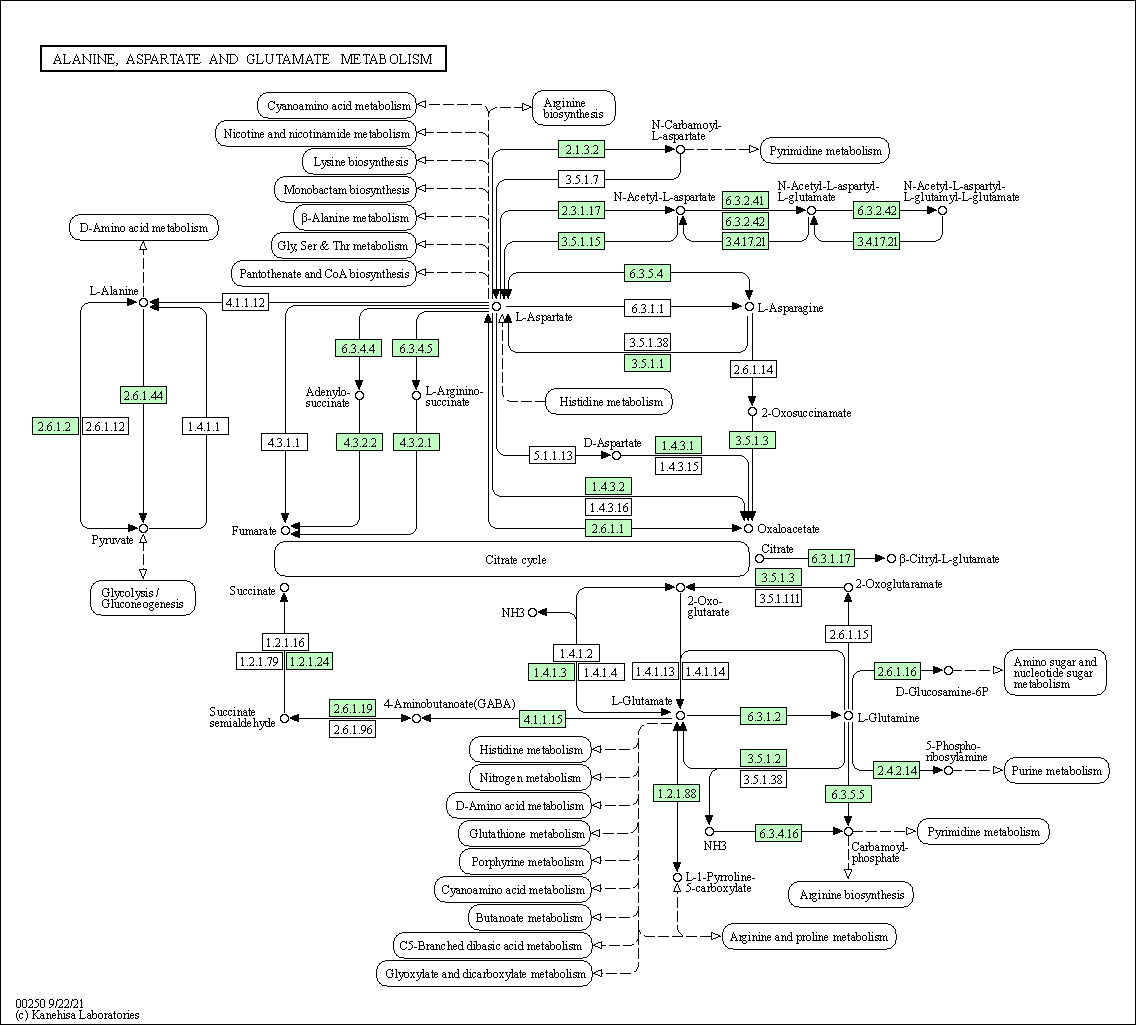
|
|||||||
| Amino sugar and nucleotide sugar metabolism | hsa00520 |
Pathway Map 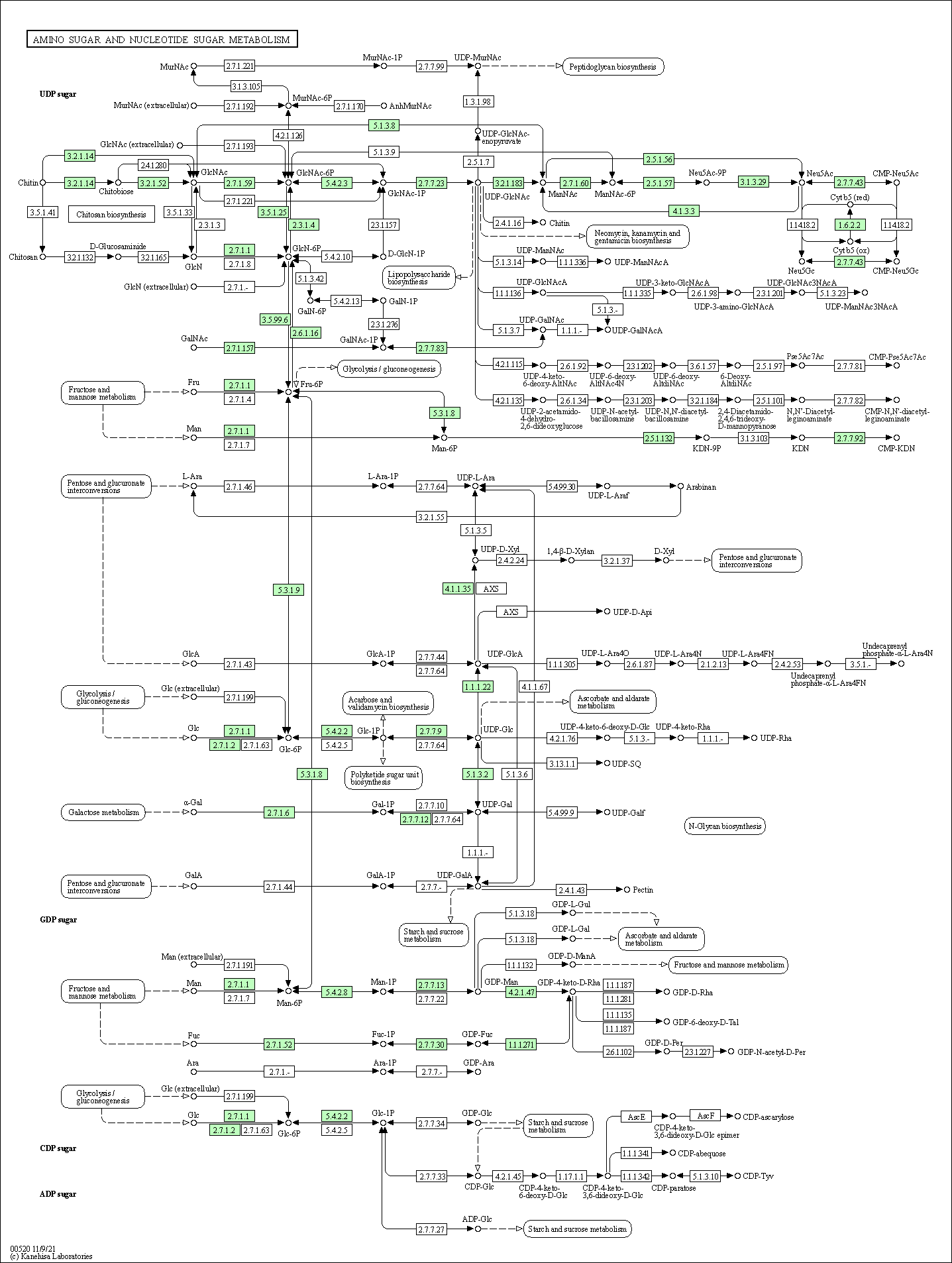
|
|||||||
| Diabetic cardiomyopathy | hsa05415 |
Pathway Map 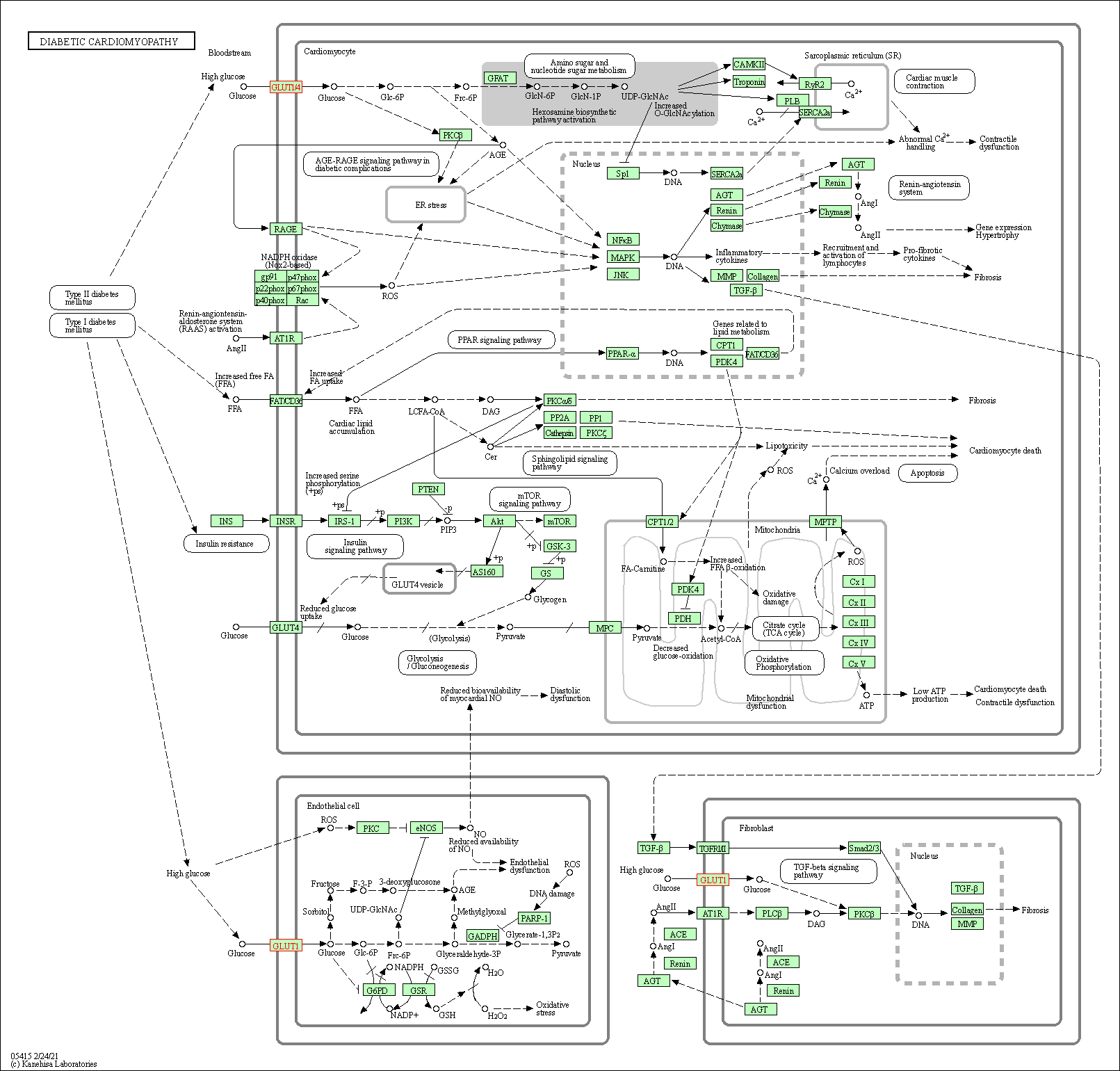
|
|||||||
| Insulin resistance | hsa04931 |
Pathway Map 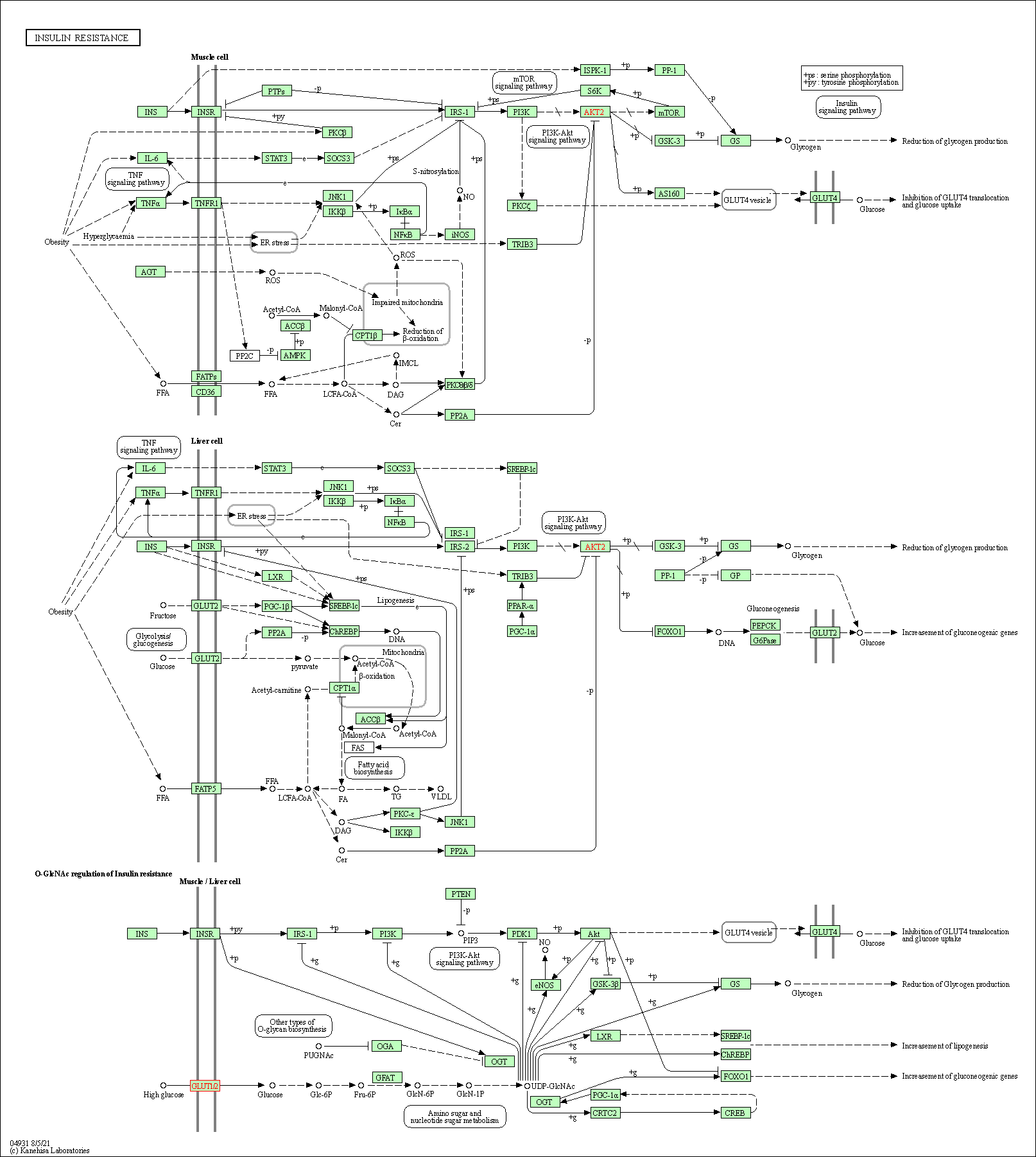
|
|||||||
| Metabolic pathways | hsa01100 |
Pathway Map 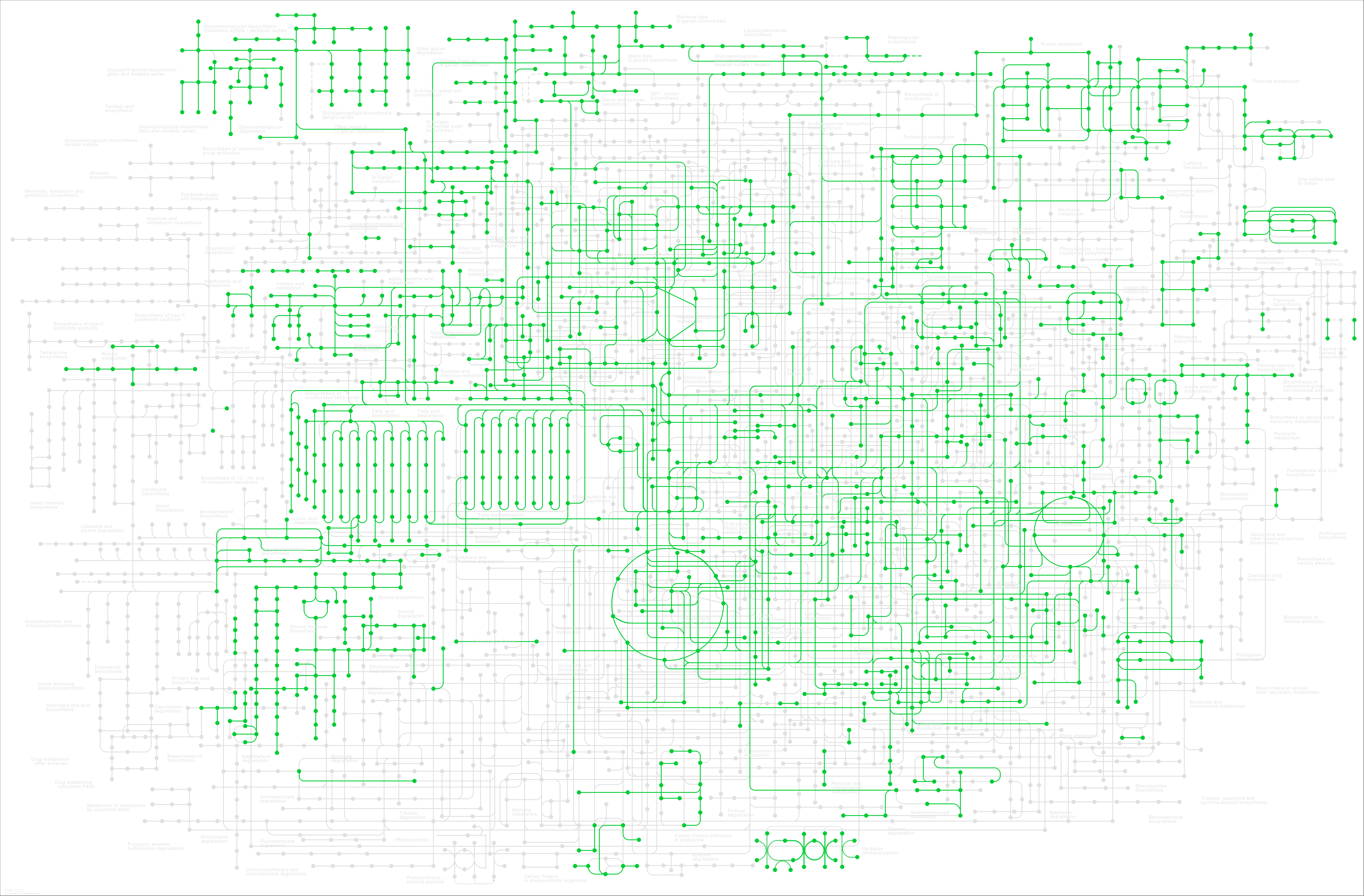
|
|||||||
| Biosynthesis of nucleotide sugars | hsa01250 |
Pathway Map 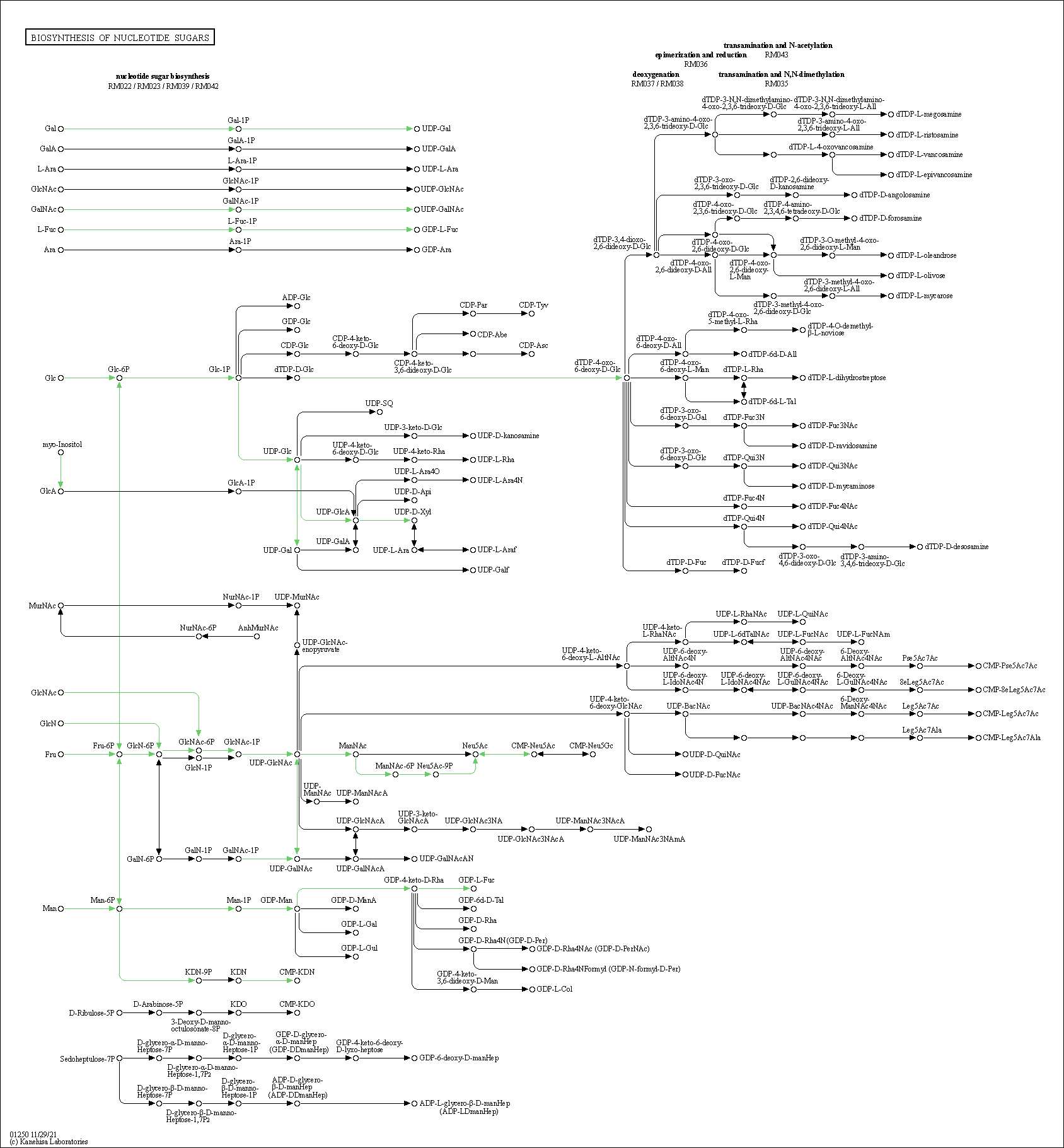
|
|||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Pro-viral | [1] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein GFPT1 in viral infection, GFPT1 protein knockout Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of GFPT1 leads to the decreased SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: ORF10 (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[2] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
ORF10
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Calu-3 cells (Human Lung Cancer Cell) (CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/USA/WA1/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[3] | |||||||
| Strains Name |
hCoV-19/USA/WA1/2020
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7.5 cells (Hepatocyte derived cellular carcinoma cell) Huh7.5 cells (Hepatocyte derived cellular carcinoma cell) (CVCL_7927 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 48 h | ||||||||
| Interaction Score | FDR ≤ 0.05 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S261
[4] |
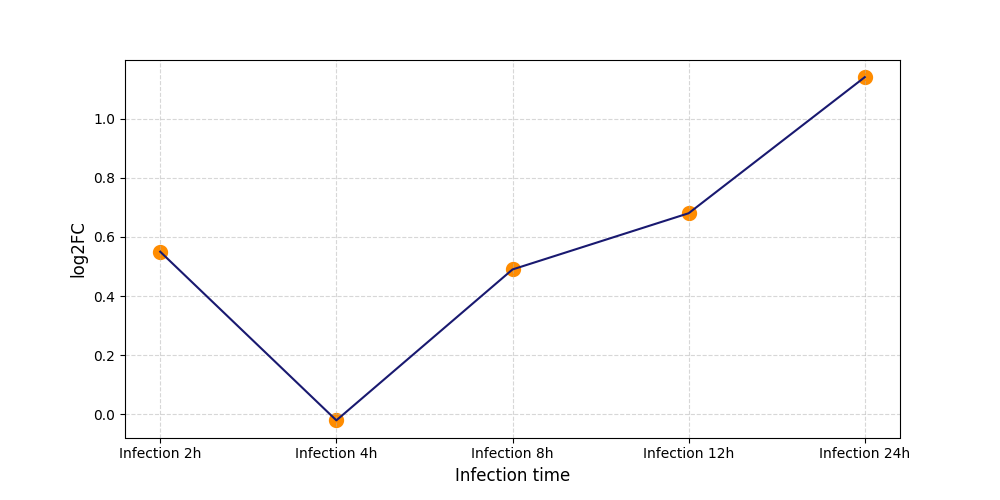 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
S261
[5] |
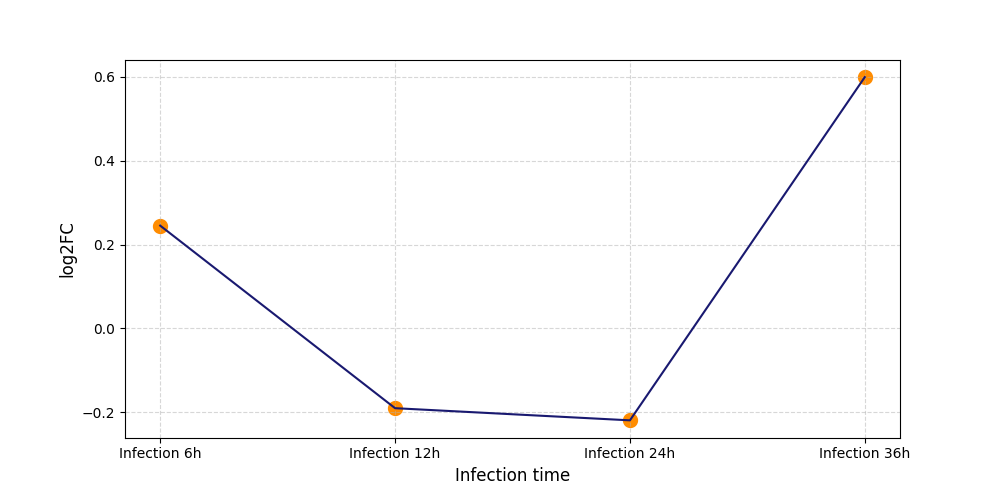 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MCGIFAYLNYHVPRTRREILETLIKGLQRLEYRGYDSAGVGFDGGNDKDWEANACKIQLIKKKGKVKALDEEVHKQQDMDLDIEFDVHLGIAHTRWATHGEPSPVNSHPQRSDKNNEFIVIHNGIITNYKDLKKFLESKGYDFESETDTETIAKLVKYMYDNRESQDTSFTTLVERVIQQLEGAFALVFKSVHFPGQAVGTRRGSPLLIGVRSEHKLSTDHIPILYRTARTQIGSKFTRWGSQGERGKDKKGSCNLSRVDSTTCLFPVEEKAVEYYFASDASAVIEHTNRVIFLEDDDVAAVVDGRLSIHRIKRTAGDHPGRAVQTLQMELQQIMKGNFSSFMQKEIFEQPESVVNTMRGRVNFDDYTVNLGGLKDHIKEIQRCRRLILIACGTSYHAGVATRQVLEELTELPVMVELASDFLDRNTPVFRDDVCFFLSQSGETADTLMGLRYCKERGALTVGITNTVGSSISRETDCGVHINAGPEIGVASTKAYTSQFVSLVMFALMMCDDRISMQERRKEIMLGLKRLPDLIKEVLSMDDEIQKLATELYHQKSVLIMGRGYHYATCLEGALKIKEITYMHSEGILAGELKHGPLALVDKLMPVIMIIMRDHTYAKCQNALQQVVARQGRPVVICDKEDTETIKNTKRTIKVPHSVDCLQGILSVIPLQLLAFHLAVLRGYDVDFPRNLAKSVTVE
Click to Show/Hide
|
|---|