Details of Host Protein
| Host Protein General Information (ID: PT1084) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
Spliceosome-associated protein 155 (SF3B1)
|
Gene Name |
SF3B1
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
SF3B1_HUMAN
|
||||||
| Protein Families |
SF3B1 family
|
||||||||
| Subcellular Location |
Nucleus speckle
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Involved in pre-mRNA splicing as a component of the splicingfactor SF3B complex. SF3B complex is required for 'A'complex assembly formed by the stable binding of U2 snRNP to thebranchpoint sequence (BPS) in pre-mRNA. Sequence independent binding ofSF3A/SF3B complex upstream of the branch site is essential, it mayanchor U2 snRNP to the pre-mRNA. Together with otherU2 snRNP complex components may also play a role in the selectiveprocessing of microRNAs (miRNAs) from the long primary miRNAtranscript, pri-miR-17-92. May also be involved in theassembly of the 'E' complex. Belongs also to theminor U12-dependent spliceosome, which is involved in the splicing ofrare class of nuclear pre-mRNA intron.
[1-3]
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| Spliceosome | hsa03040 |
Pathway Map 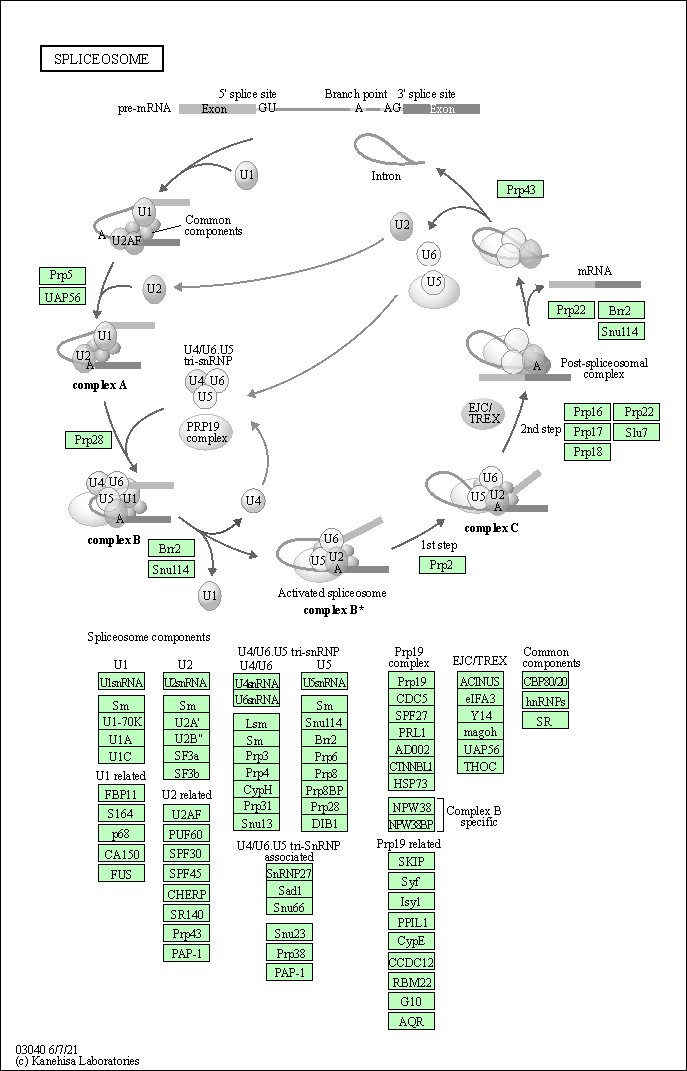
|
|||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Anti-viral | [4] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein SF3B1 in viral infection, SF3B1 protein Knockdown Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of SF3B1 increases SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: ORF10 (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
ORF10
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (human liver cell line) (CVCL_0336 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| RNA Region: E region (hCoV-19/Zambia/ZMB-88673/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Zambia/ZMB-88673/2021
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Malawi/KRISP-K010154/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Malawi/KRISP-K010154/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: E region (hCoV-19/Eswatini/N2779/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Eswatini/N2779/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Wuhan/WHUH020/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan/WHUH020/2020
|
||||||||
| Strains Family |
Epsilon (B.1.427; B.1.429)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Wuhan/HBCDC-HB-04/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan/HBCDC-HB-04/2020
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/USA/CA-CZB-14678/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/USA/CA-CZB-14678/2020
|
||||||||
| Strains Family |
Epsilon (B.1.427; B.1.429)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Philippines/PH-PGC-104159/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Philippines/PH-PGC-104159/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/NewZealand/20CV0676/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/NewZealand/20CV0676/2020
|
||||||||
| Strains Family |
Epsilon (B.1.427; B.1.429)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Myanmar/DSMRC-065/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Myanmar/DSMRC-065/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Malaysia/MGI_GS0622/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Malaysia/MGI_GS0622/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Guam/GU-CDC-2-3906081-/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Guam/GU-CDC-2-3906081-/2021
|
||||||||
| Strains Family |
Epsilon (B.1.427; B.1.429)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Ethiopia/ILRI_COVM02835/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Ethiopia/ILRI_COVM02835/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/ElSalvador/INC-LNSP-153/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/ElSalvador/INC-LNSP-153/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Chile/AT-59206/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Chile/AT-59206/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Canada/UN-276648/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Canada/UN-276648/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Bonaire/BQ-RIVM-101067/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Bonaire/BQ-RIVM-101067/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Bangladesh/CHRF-GSB17-117/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Bangladesh/CHRF-GSB17-117/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: M region (hCoV-19/Australia/SA60463/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Australia/SA60463/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-SARS/Manitoba/2003 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-SARS/Manitoba/2003
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome-related coronavirus (HCoV-SARS)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-OC43/VR-759 Quebec/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-OC43/VR-759 Quebec/2019
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus OC43 (HCov-OC43)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-NL63/Amsterdam/2004 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-NL63/Amsterdam/2004
|
||||||||
| Strains Family |
Alpha
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human Coronavirus NL63 (HCov-NL63)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-MERS/Jeddah/2012 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-MERS/Jeddah/2012
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Middle East respiratory syndrome-related coronavirus (HCov-MERS)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-HKU1/Hong Kong/2004 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-HKU1/Hong Kong/2004
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus HKU1 (HCoV-HKU1)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-229E/Würzburg/2000 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-229E/Würzburg/2000
|
||||||||
| Strains Family |
Alpha
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus 229E (HCov-229E)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
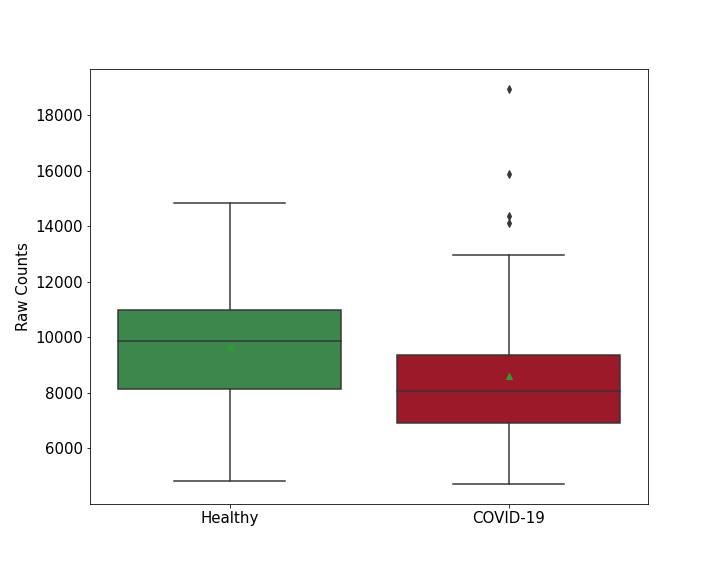
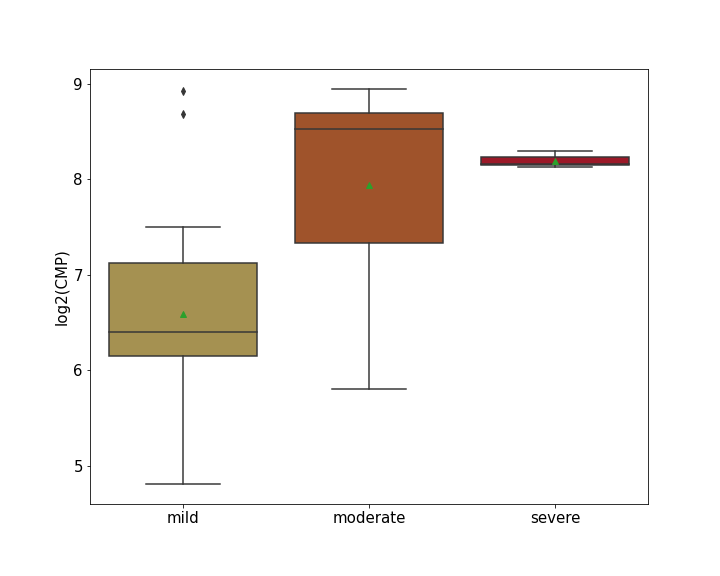
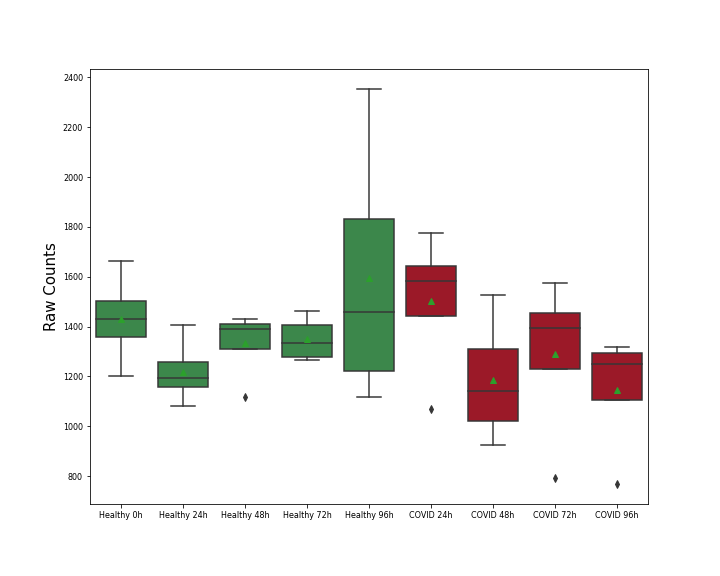
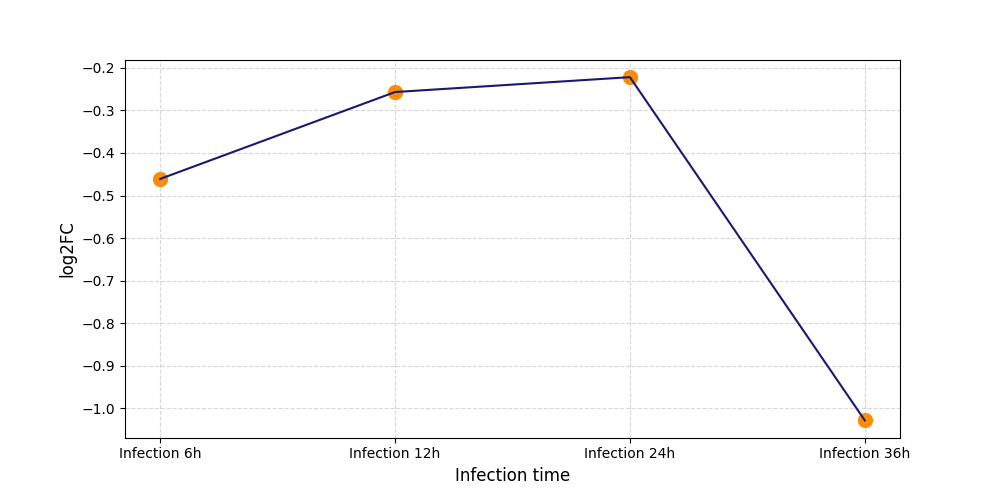
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
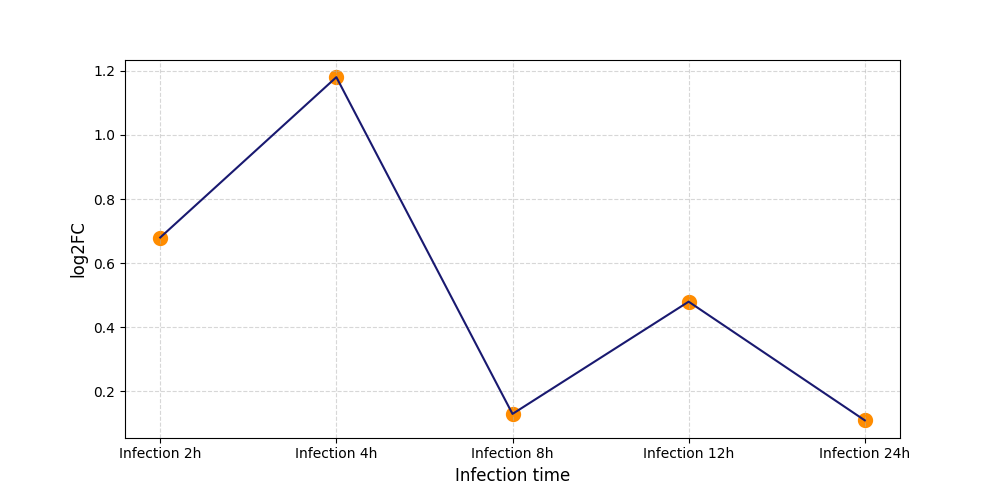
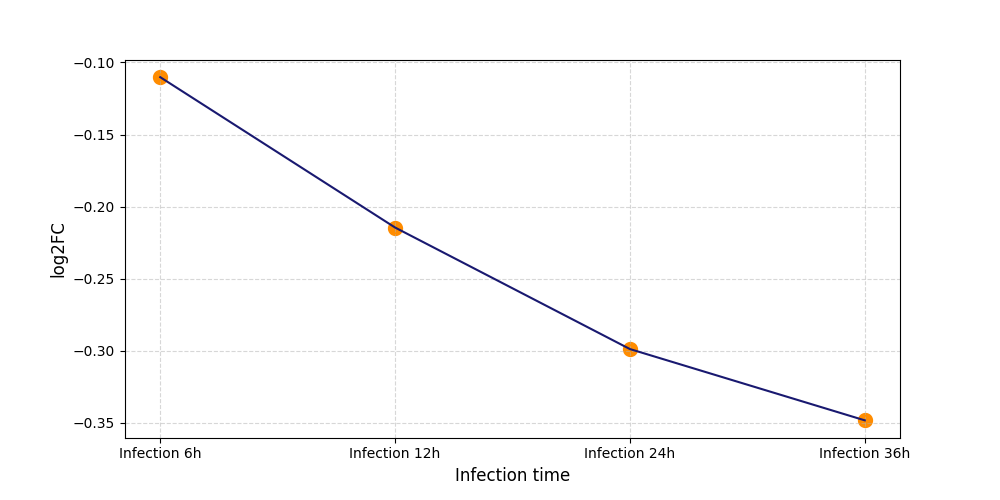
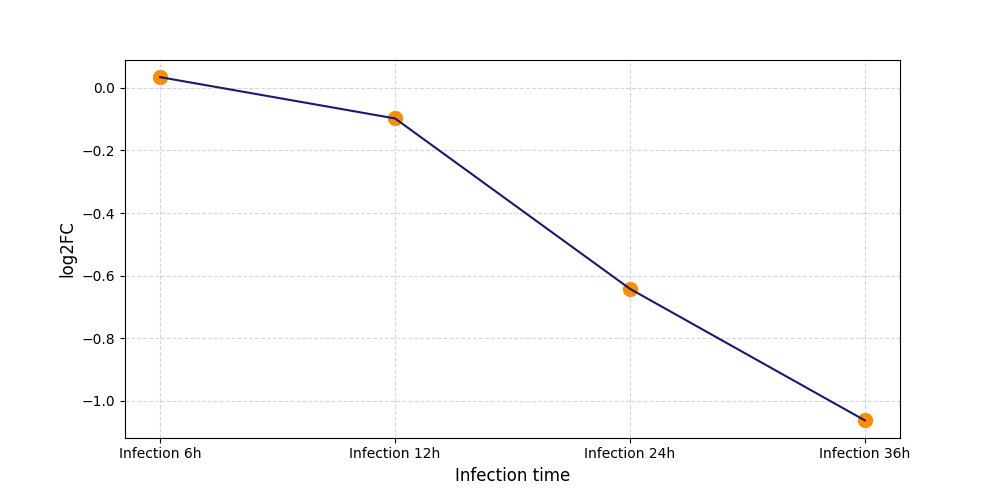
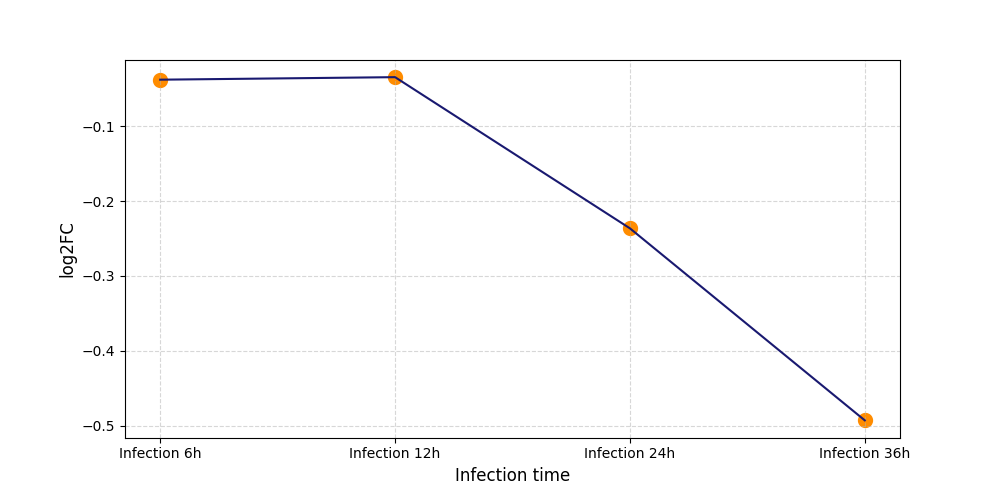
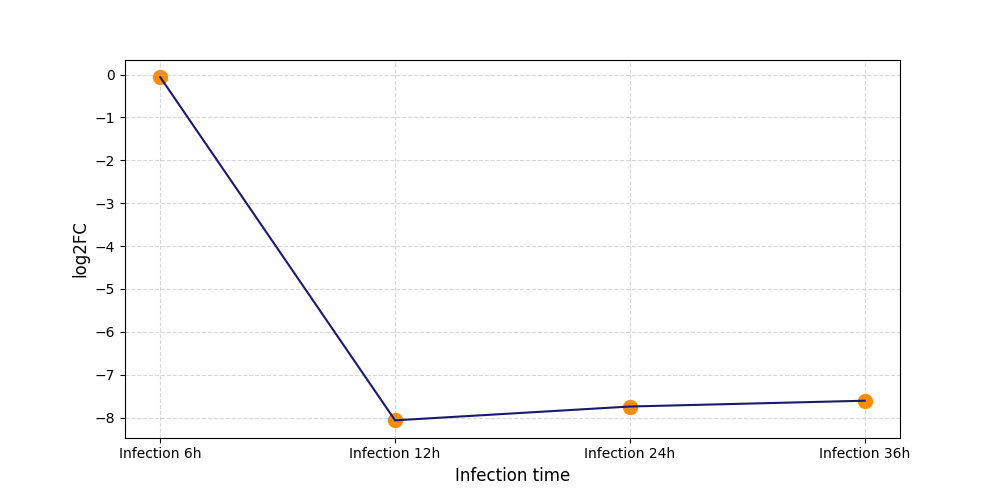
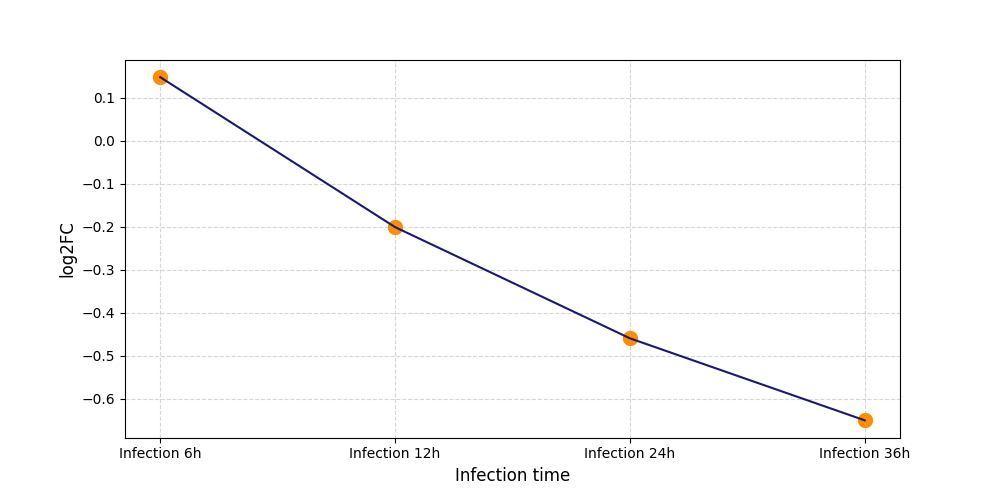
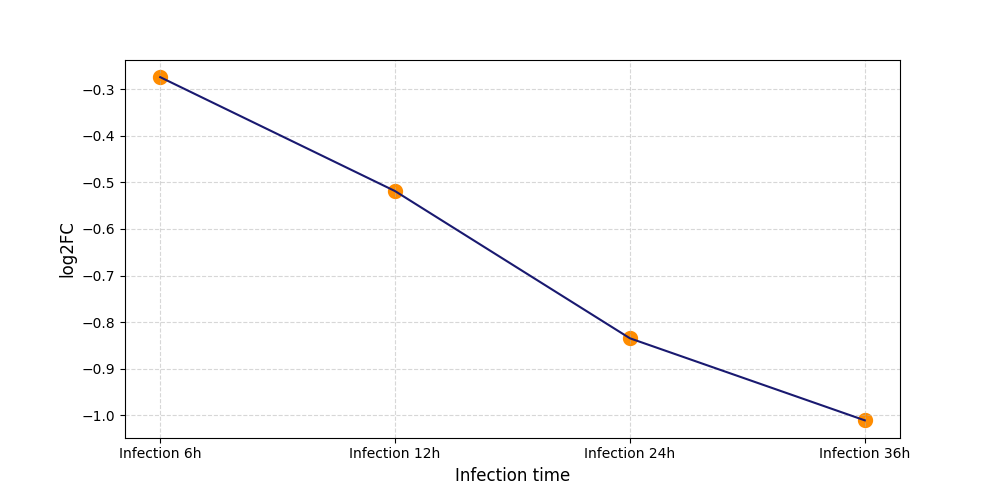
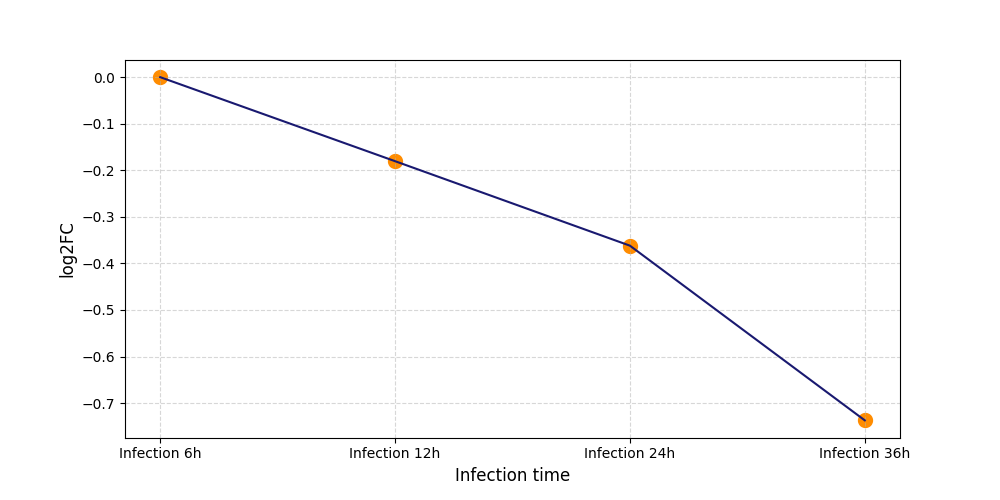
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S129
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
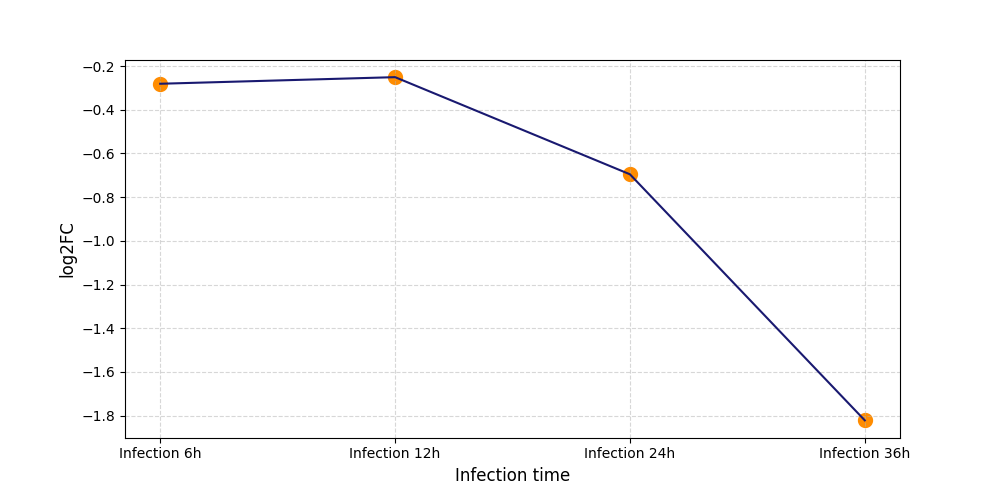
S129
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
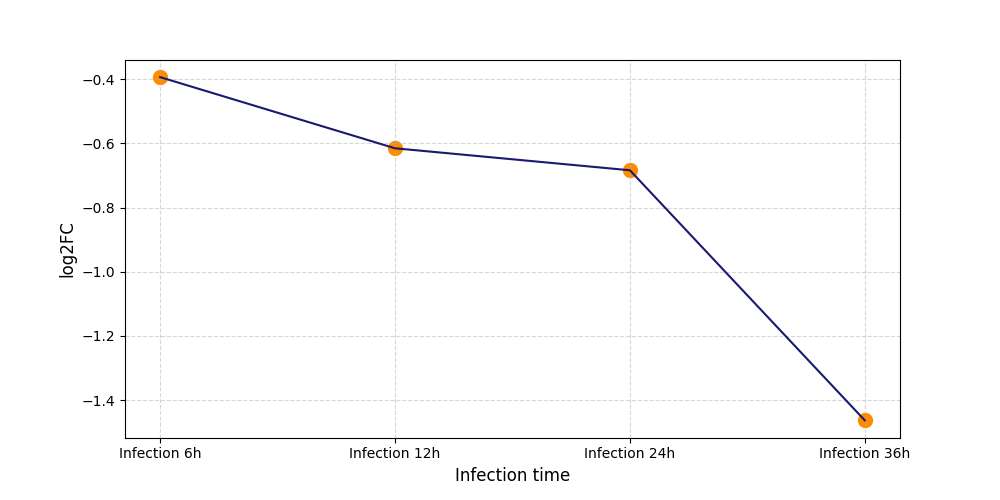
S194
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
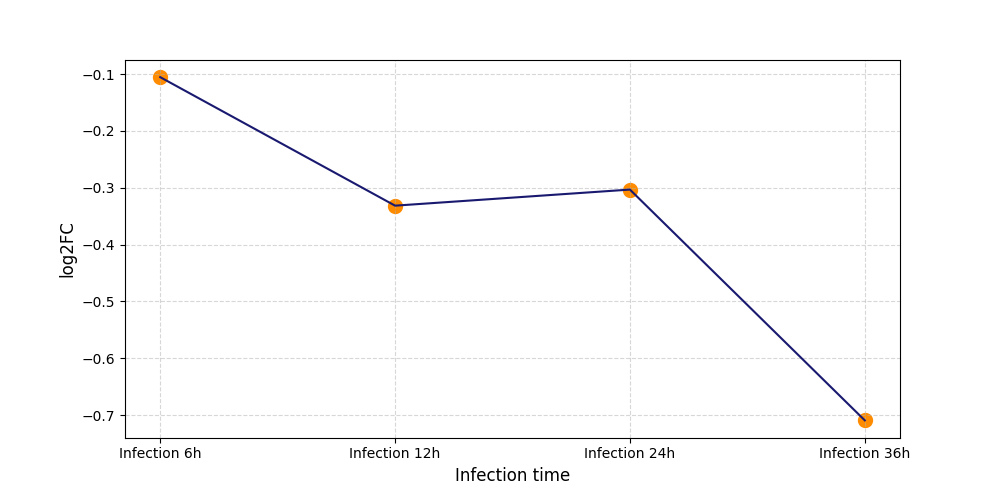
S308
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
S332
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
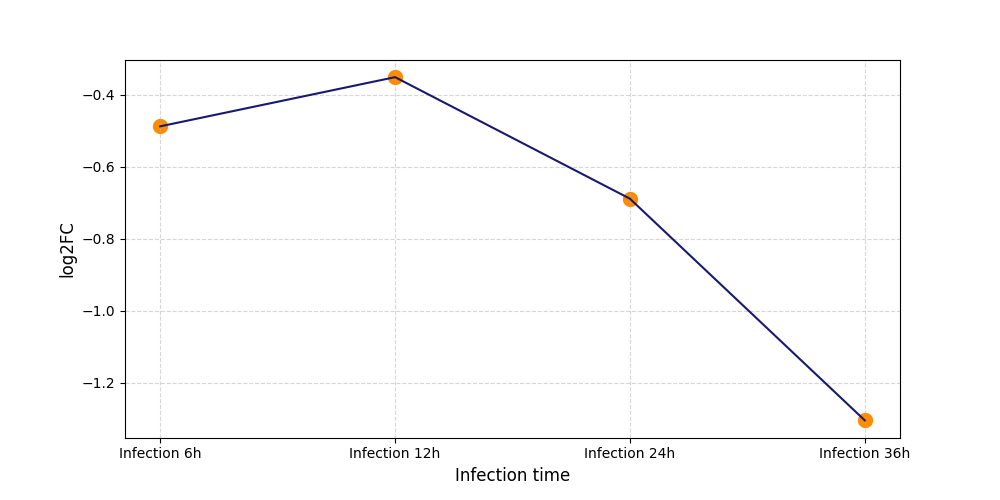
|
T142
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
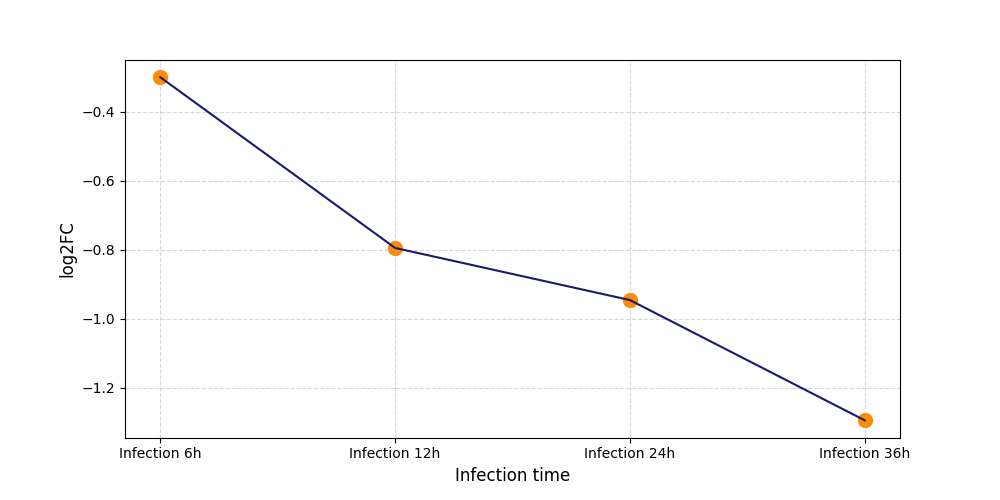
|
T211
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
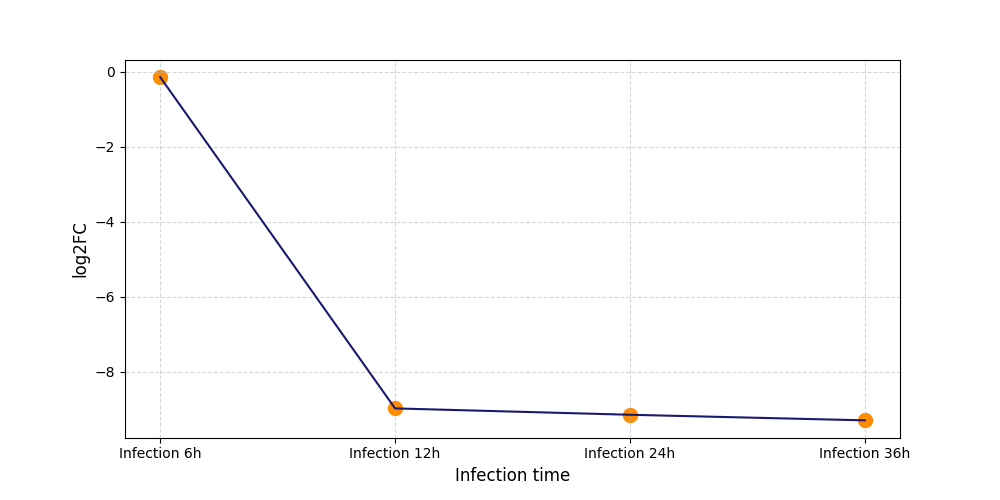
|
T223
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T227
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
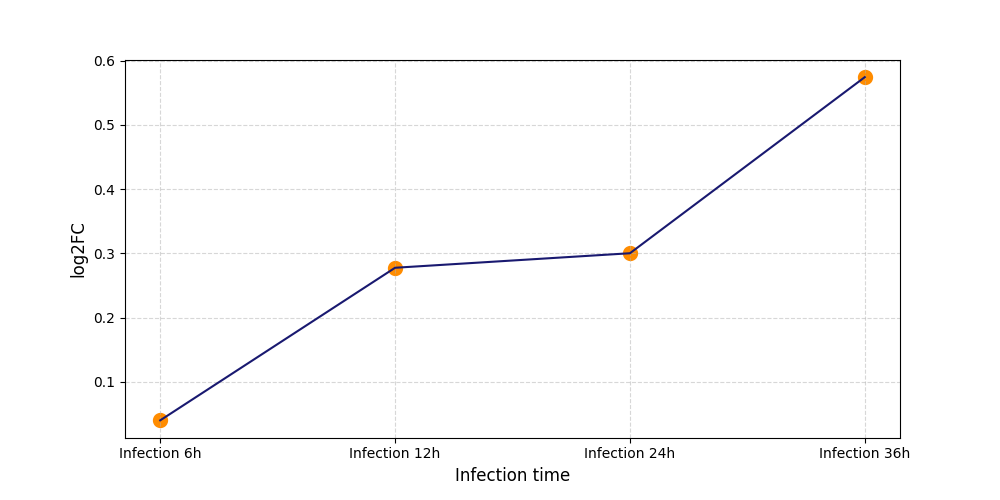
T235
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
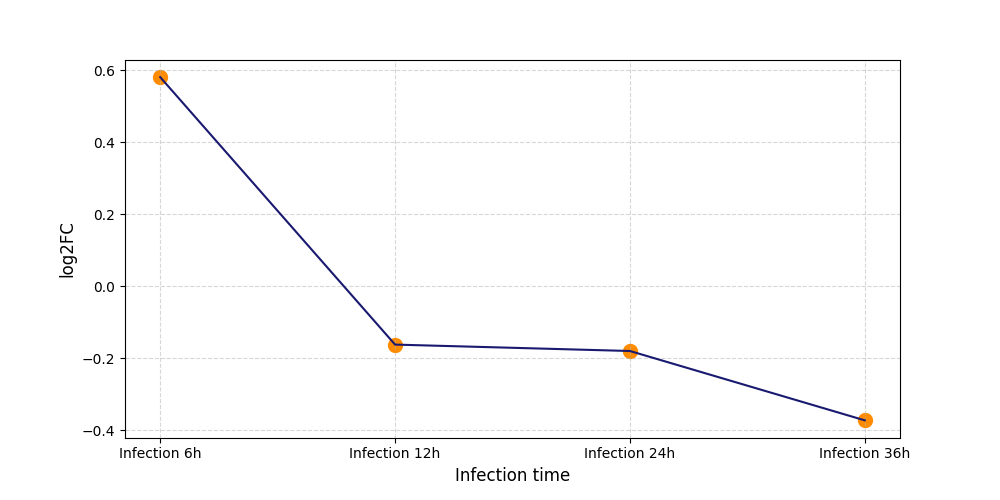
T244
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
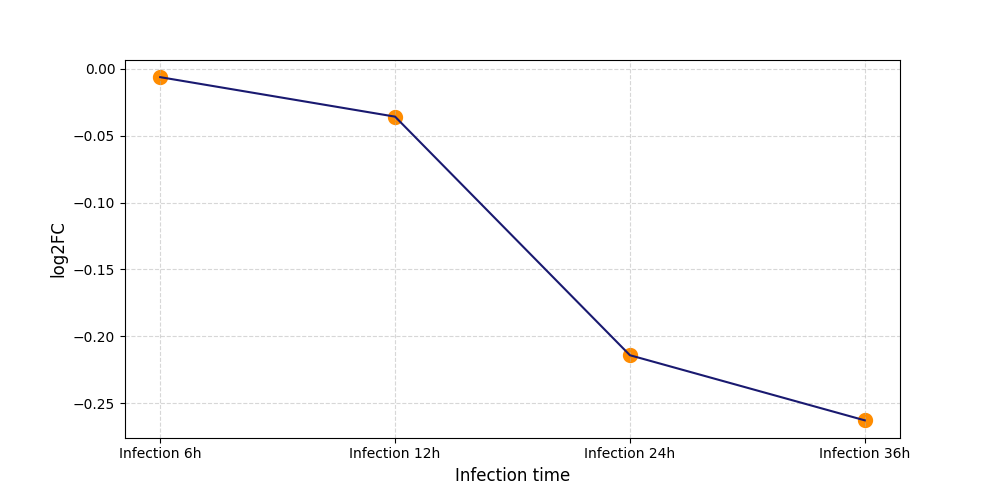
T248
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T257
[8] |
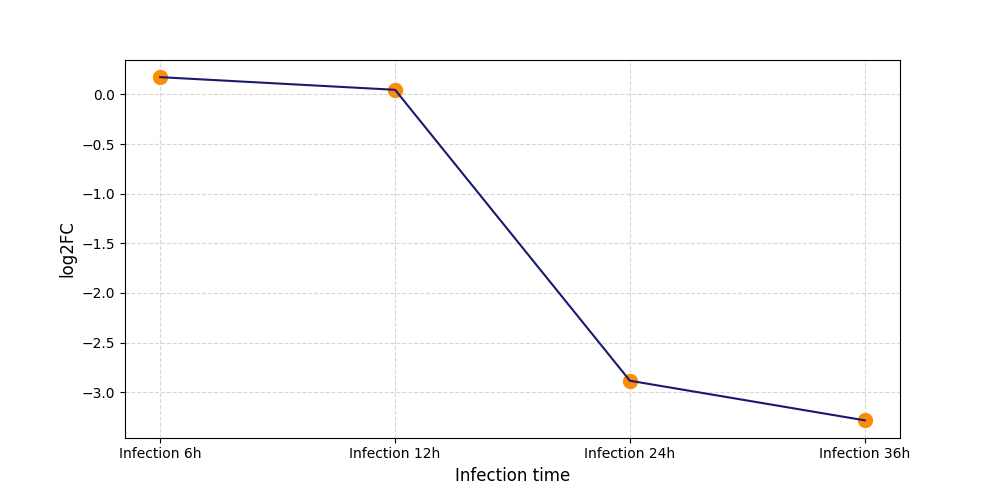 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T261
[8] |
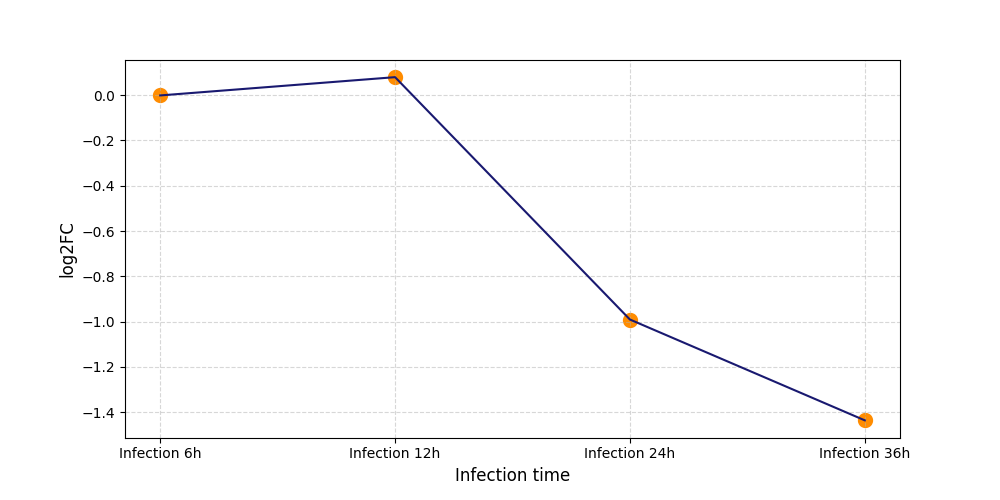 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T267
[8] |
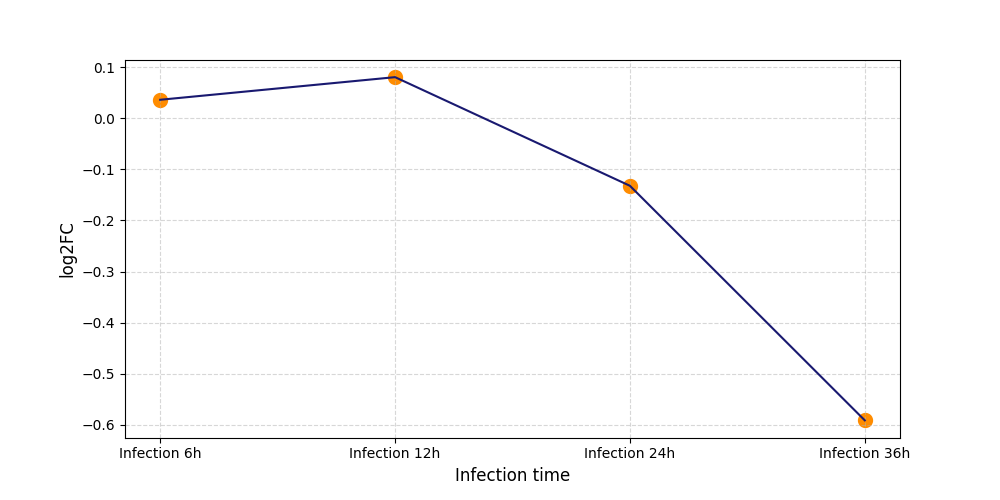 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T273
[8] |
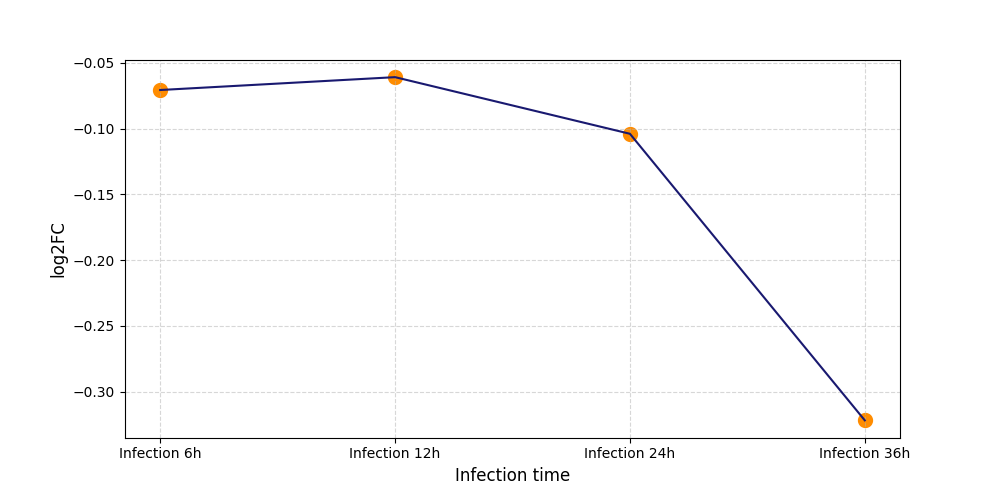 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T278
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T303
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T313
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T326
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T328
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
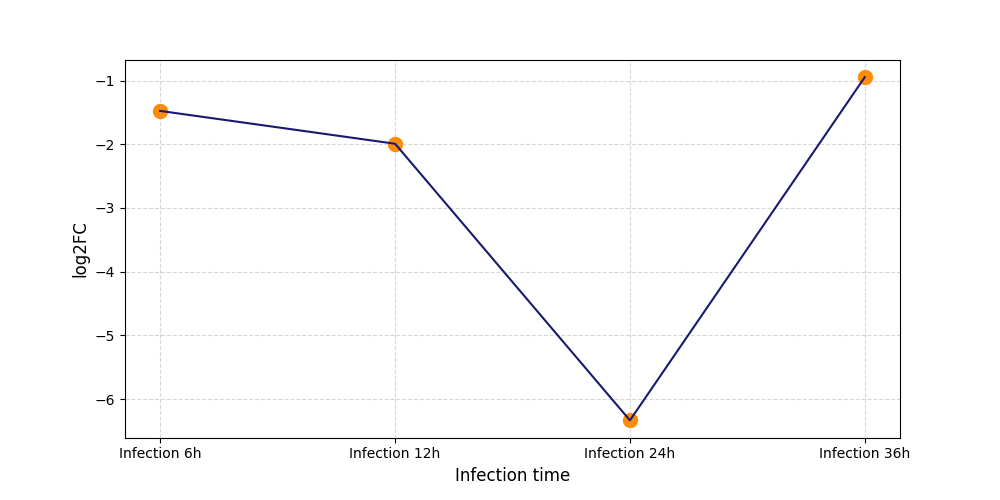
|
T350
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
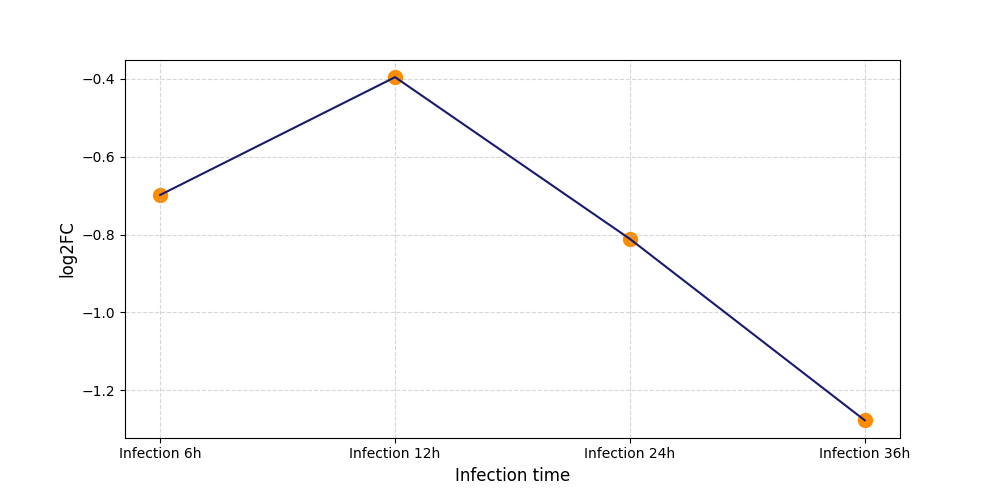
|
T426
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
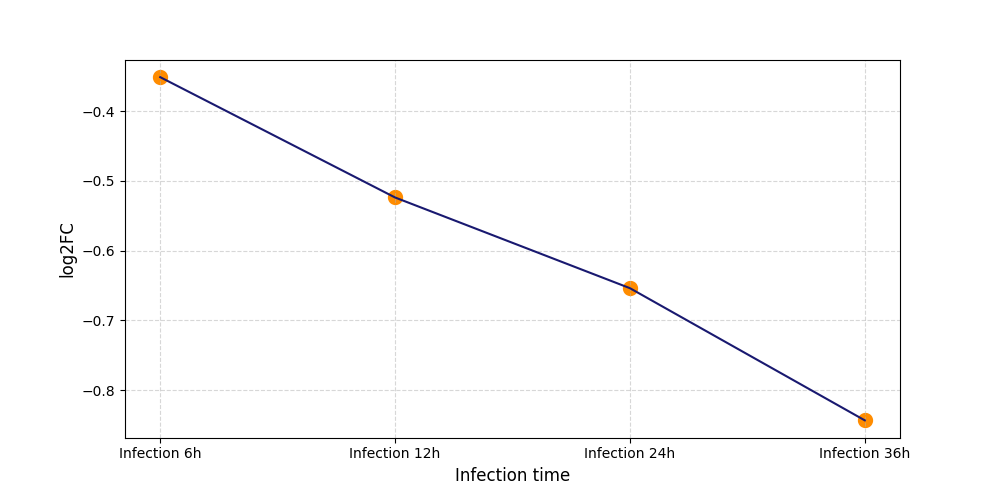
|
T432
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Y421
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MAKIAKTHEDIEAQIREIQGKKAALDEAQGVGLDSTGYYDQEIYGGSDSRFAGYVTSIAATELEDDDDDYSSSTSLLGQKKPGYHAPVALLNDIPQSTEQYDPFAEHRPPKIADREDEYKKHRRTMIISPERLDPFADGGKTPDPKMNARTYMDVMREQHLTKEEREIRQQLAEKAKAGELKVVNGAAASQPPSKRKRRWDQTADQTPGATPKKLSSWDQAETPGHTPSLRWDETPGRAKGSETPGATPGSKIWDPTPSHTPAGAATPGRGDTPGHATPGHGGATSSARKNRWDETPKTERDTPGHGSGWAETPRTDRGGDSIGETPTPGASKRKSRWDETPASQMGGSTPVLTPGKTPIGTPAMNMATPTPGHIMSMTPEQLQAWRWEREIDERNRPLSDEELDAMFPEGYKVLPPPAGYVPIRTPARKLTATPTPLGGMTGFHMQTEDRTMKSVNDQPSGNLPFLKPDDIQYFDKLLVDVDESTLSPEEQKERKIMKLLLKIKNGTPPMRKAALRQITDKAREFGAGPLFNQILPLLMSPTLEDQERHLLVKVIDRILYKLDDLVRPYVHKILVVIEPLLIDEDYYARVEGREIISNLAKAAGLATMISTMRPDIDNMDEYVRNTTARAFAVVASALGIPSLLPFLKAVCKSKKSWQARHTGIKIVQQIAILMGCAILPHLRSLVEIIEHGLVDEQQKVRTISALAIAALAEAATPYGIESFDSVLKPLWKGIRQHRGKGLAAFLKAIGYLIPLMDAEYANYYTREVMLILIREFQSPDEEMKKIVLKVVKQCCGTDGVEANYIKTEILPPFFKHFWQHRMALDRRNYRQLVDTTVELANKVGAAEIISRIVDDLKDEAEQYRKMVMETIEKIMGNLGAADIDHKLEEQLIDGILYAFQEQTTEDSVMLNGFGTVVNALGKRVKPYLPQICGTVLWRLNNKSAKVRQQAADLISRTAVVMKTCQEEKLMGHLGVVLYEYLGEEYPEVLGSILGALKAIVNVIGMHKMTPPIKDLLPRLTPILKNRHEKVQENCIDLVGRIADRGAEYVSAREWMRICFELLELLKAHKKAIRRATVNTFGYIAKAIGPHDVLATLLNNLKVQERQNRVCTTVAIAIVAETCSPFTVLPALMNEYRVPELNVQNGVLKSLSFLFEYIGEMGKDYIYAVTPLLEDALMDRDLVHRQTASAVVQHMSLGVYGFGCEDSLNHLLNYVWPNVFETSPHVIQAVMGALEGLRVAIGPCRMLQYCLQGLFHPARKVRDVYWKIYNSIYIGSQDALIAHYPRIYNDDKNTYIRYELDYIL
Click to Show/Hide
|
|---|