Details of Host Protein
| Host Protein General Information (ID: PT0364) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
Double-stranded RNA-specific adenosine deaminase (ADAR)
|
Gene Name |
ADAR
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
DSRAD_HUMAN
|
||||||
| EC Number |
3.5.4.37
|
||||||||
| Subcellular Location |
Cytoplasm Nucleus
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Catalyzes the hydrolytic deamination of adenosine to inosinein double-stranded RNA (dsRNA) referred to as A-to-I RNA editing. This may affect geneexpression and function in a number of ways that include mRNAtranslation by changing codons and hence the amino acid sequence ofproteins since the translational machinery read the inosine as aguanosine; pre-mRNA splicing by altering splice site recognitionsequences; RNA stability by changing sequences involved in nucleaserecognition; genetic stability in the case of RNA virus genomes bychanging sequences during viral RNA replication; and RNA structure-dependent activities such as microRNA production or targeting orprotein-RNA interactions. Can edit both viral and cellular RNAs and canedit RNAs at multiple sites (hyper-editing) or at specific sites (site-specific editing). Its cellular RNA substrates include: bladder cancer-associated protein (BLCAP), neurotransmitter receptors for glutamate (GRIA2) and serotonin (HTR2C) and GABA receptor (GABRA3). Site-specificRNA editing of transcripts encoding these proteins results in aminoacid substitutions which consequently alters their functionalactivities. Exhibits low-level editing at the GRIA2 Q/R site, but editsefficiently at the R/G site and HOTSPOT1. Its viral RNA substratesinclude: hepatitis C virus (HCV), vesicular stomatitis virus (VSV), measles virus (MV), hepatitis delta virus (HDV), and humanimmunodeficiency virus type 1 (HIV-1). Exhibits either a proviral (HDV, MV, VSV and HIV-1) or an antiviral effect (HCV) and this can beediting-dependent (HDV and HCV), editing-independent (VSV and MV) orboth (HIV-1). Impairs HCV replication via RNA editing at multiplesites. Enhances the replication of MV, VSV and HIV-1 through anediting-independent mechanism via suppression of EIF2AK2/PKR activationand function. Stimulates both the release and infectivity of HIV-1viral particles by an editing-dependent mechanism where it associateswith viral RNAs and edits adenosines in the 5'UTR and the Rev and Tatcoding sequence. Can enhance viral replication of HDV via A-to-Iediting at a site designated as amber/W, thereby changing an UAG amberstop codon to an UIG tryptophan (W) codon that permits synthesis of thelarge delta antigen (L-HDAg) which has a key role in the assembly ofviral particles. However, high levels of ADAR1 inhibit HDV replication.
[1-14]
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| Coronavirus disease - COVID-19 | hsa05171 |
Pathway Map 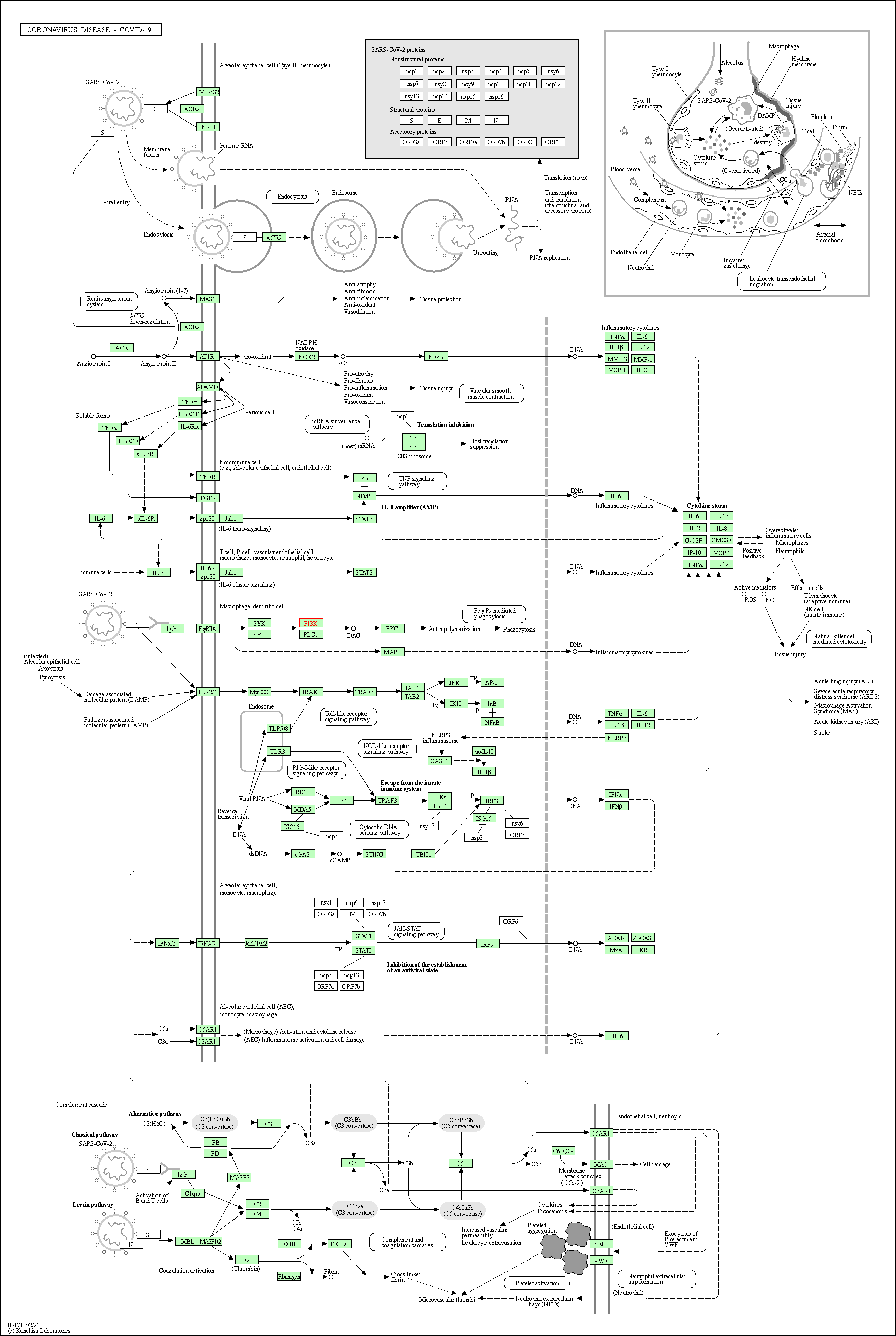
|
|||||||
| Cytosolic DNA-sensing pathway | hsa04623 |
Pathway Map 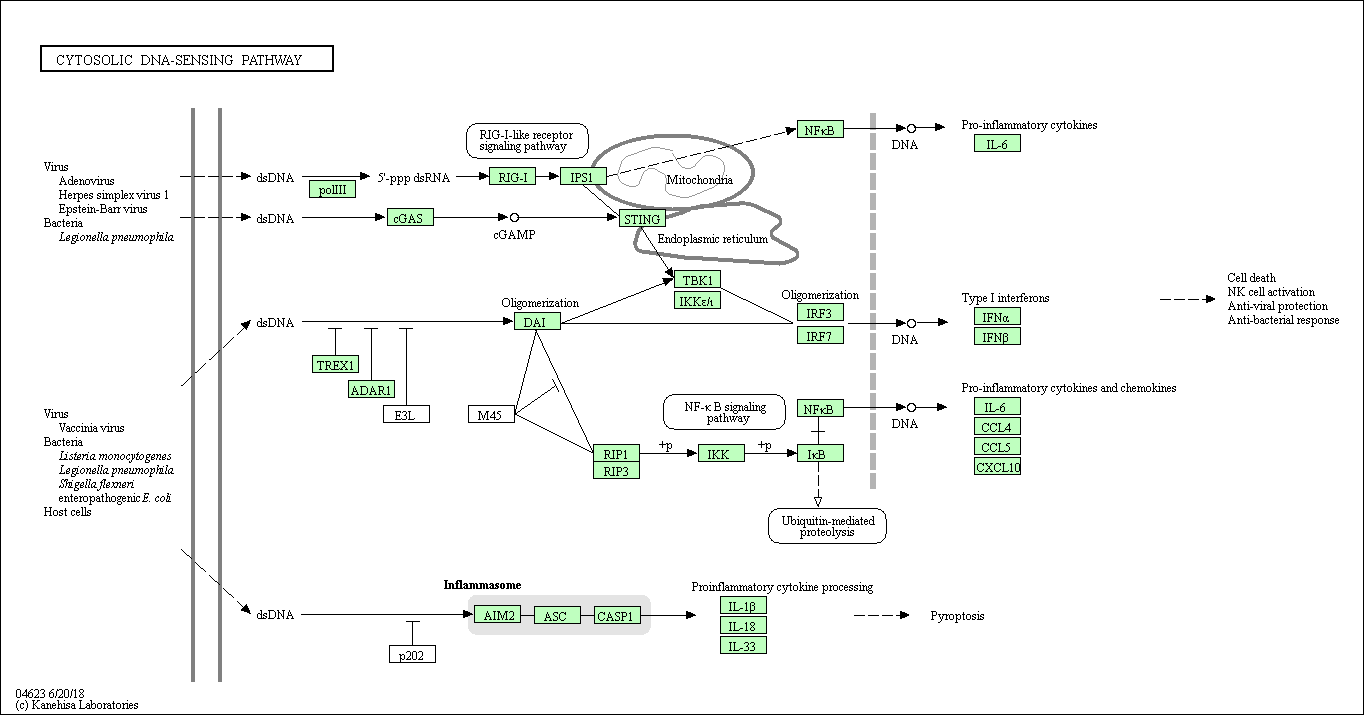
|
|||||||
| Measles | hsa05162 |
Pathway Map 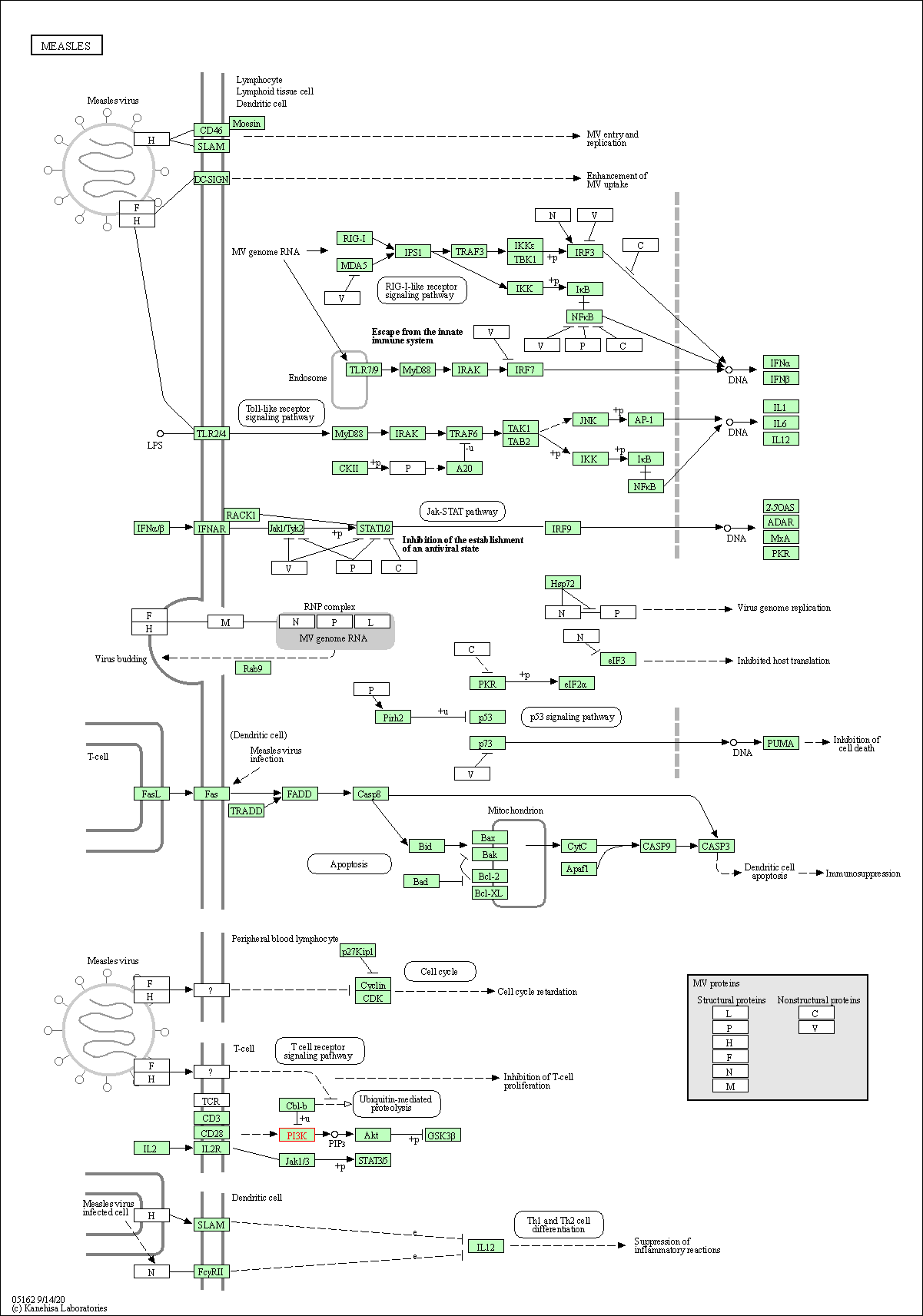
|
|||||||
| Influenza A | hsa05164 |
Pathway Map 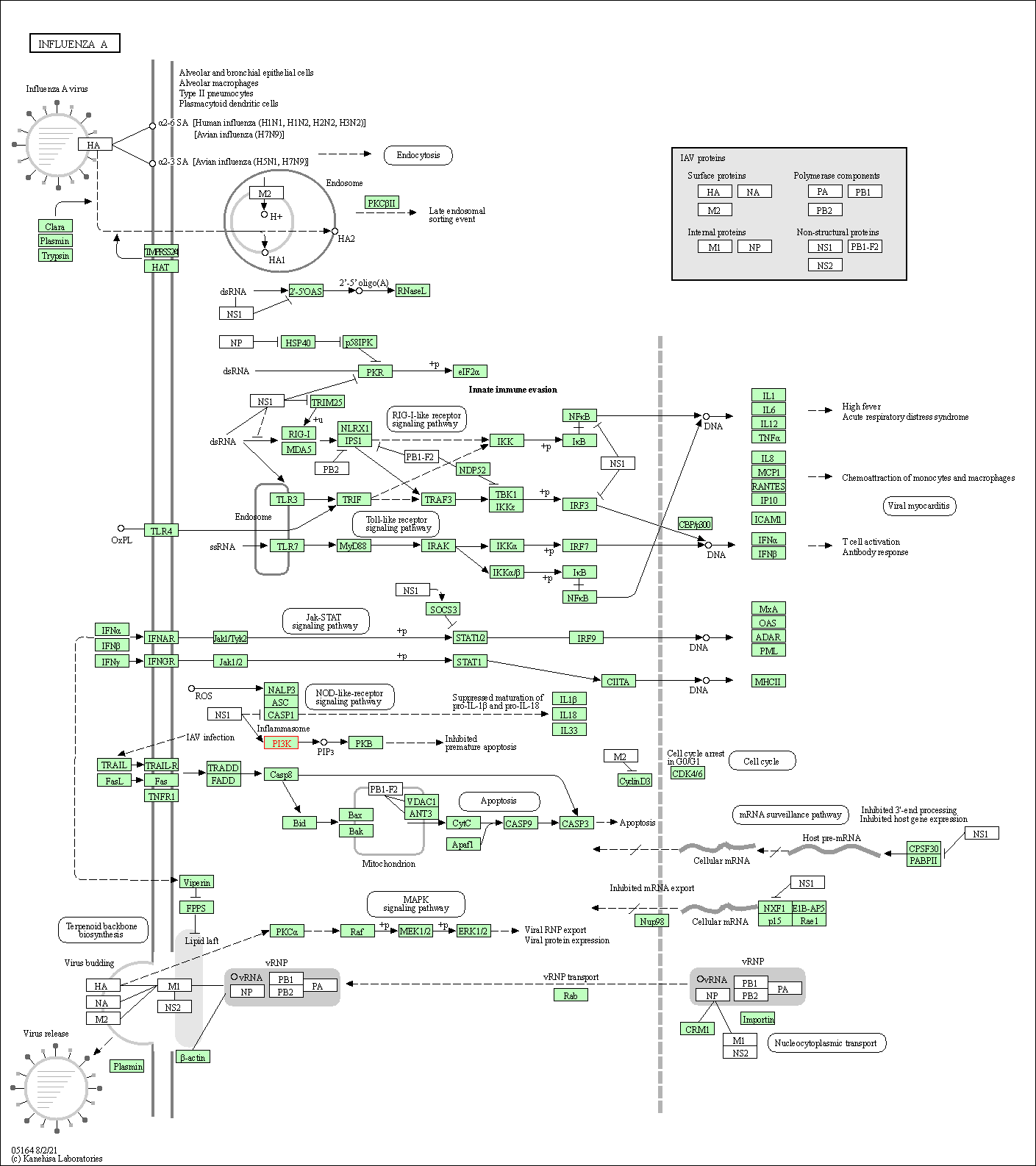
|
|||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Pro-viral | [15] | |||||
| Infected Tissue | Lung | Infection Time | 24 h | ||||||
| Infected Cell | A549 Cells (Adenocarcinomic Human alveolar basal epithelial cells) | Cellosaurus ID | CVCL_H249 | ||||||
| Method Description | To detect the role of host protein ADAR in viral infection, ADAR protein knockout A549 Cells were infected with SARS-COV-2 for 24 h , and the effects on infection was detected through qRT-PCR. | ||||||||
| Results | It is reported that Knockdown of ADAR leads to the decreased the 50% percent of infected cells compared with control group. | ||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: 3'-UTR (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[16] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) (CVCL_0336;CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Liver; Lung | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| RNA Region: ORF10 (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[16] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF10
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) (CVCL_0336;CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Liver; Lung | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| RNA Region: 5'-UTR-L (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[16] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
5'-UTR-L
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) (CVCL_0336;CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Liver; Lung | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[17] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (liver carcinoma cell) Huh7 cells (liver carcinoma cell) (CVCL_0336 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
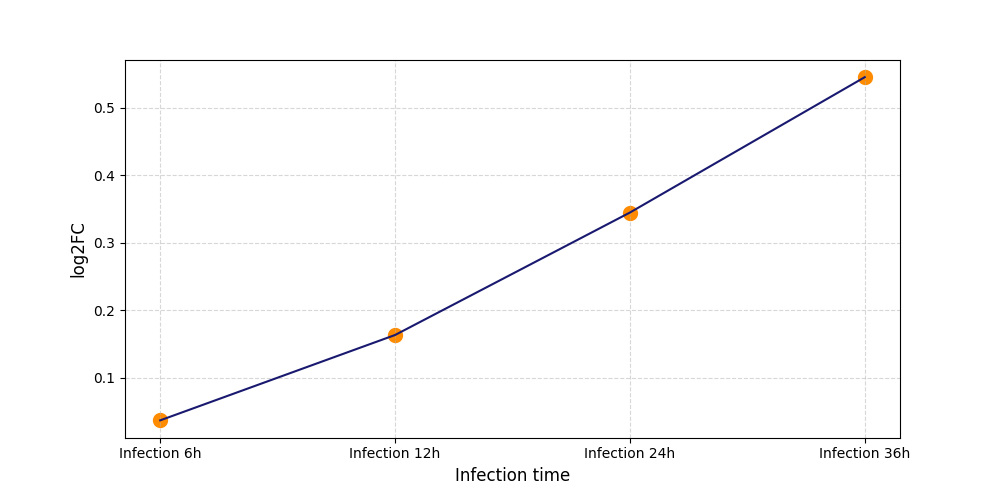
| Infection Time | 24h | ||||||||
| Interaction Score | log2FC = 1.19537E+14 | ||||||||
| Method Description | RNA antisense purification and quantitative mass spectrometry (RAP-MS); Tandem mass tag (TMT) labelling; liquid chromatography tandem mass spectrometry (LC-MS/MS); Westernblot | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/France/IDF-220-95/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[15] | |||||||
| Strains Name |
hCoV-19/France/IDF-220-95/2020
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Direct interaction | ||||||||
| Infection Cells | HEK293 Cells (Human embryonic kidney cell) HEK293 Cells (Human embryonic kidney cell) (CVCL_0045 ) | ||||||||
| Cell Originated Tissue | Kidney | ||||||||
| Infection Time | 48 h | ||||||||
| Interaction Score | SAINT score ≥ 0.79 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/England/02/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[18] | |||||||
| Strains Name |
hCoV-19/England/02/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Calu-3 cells (Human lung cancer cell) Calu-3 cells (Human lung cancer cell) (CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Lung | ||||||||
| Infection Time | 24 h | ||||||||
| Interaction Score | P-adjust = 0.036 | ||||||||
| Method Description | UV protein-RNA crosslinking; RNA interactome capture (cRIC); RNA antisense purification coupled with mass spectrometry (RAP-MS) | ||||||||
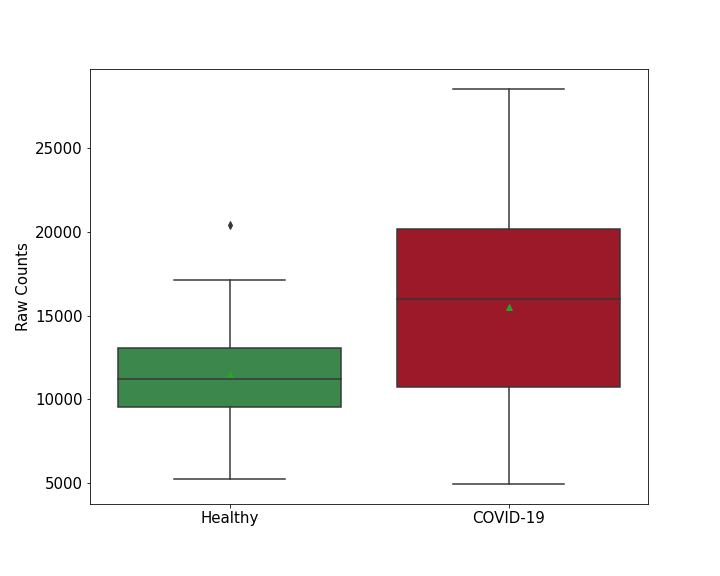
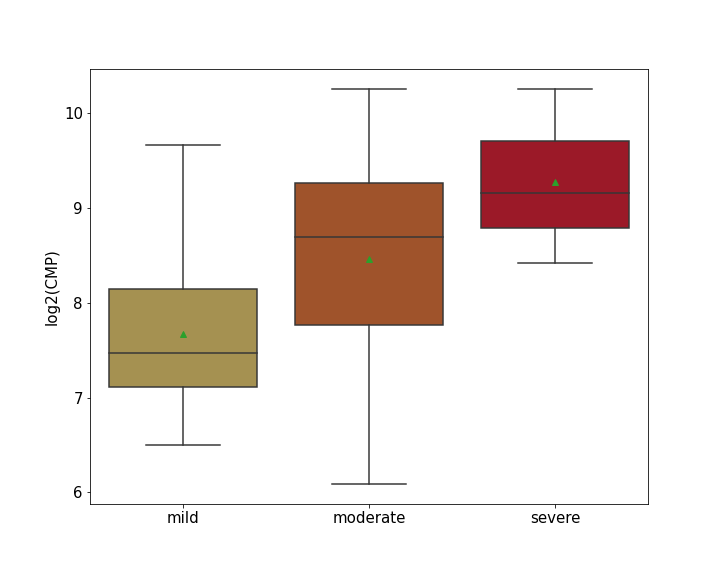
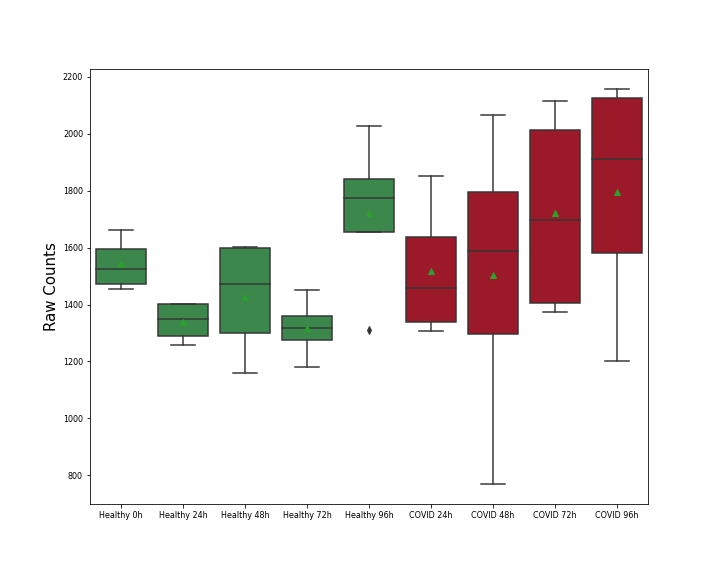
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S599
[19] |
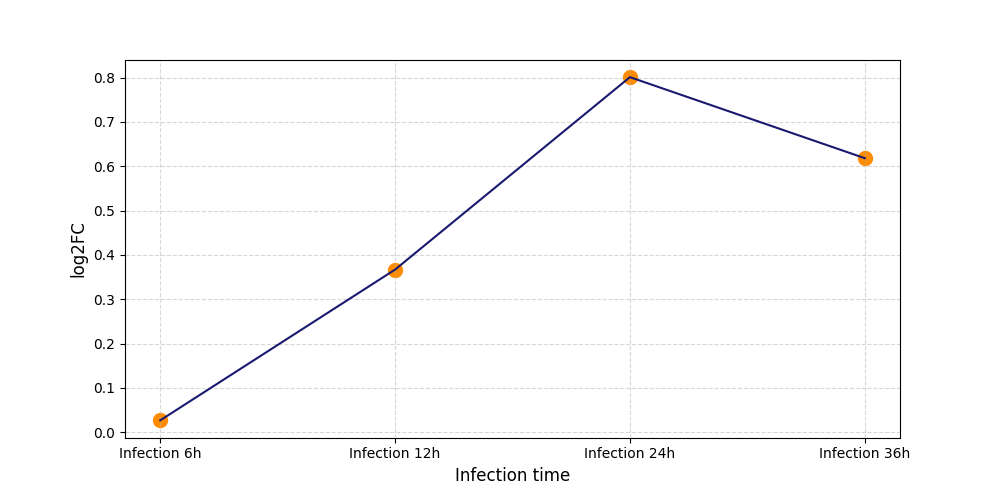 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
S614
[19] |
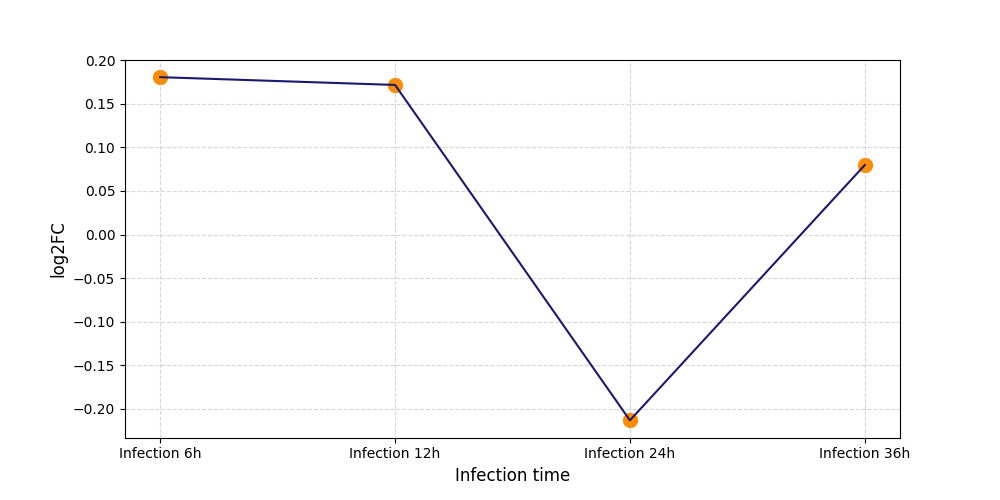 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
S825
[19] |
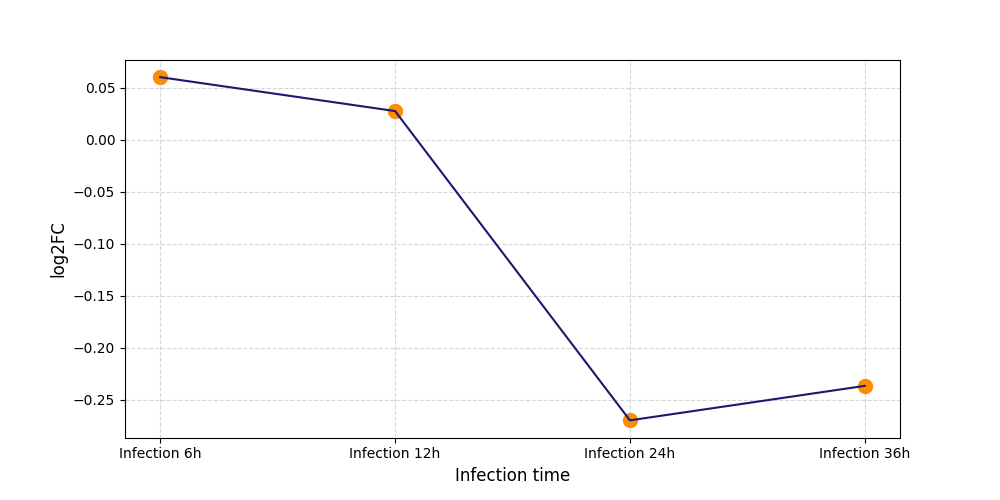 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T601
[19] |
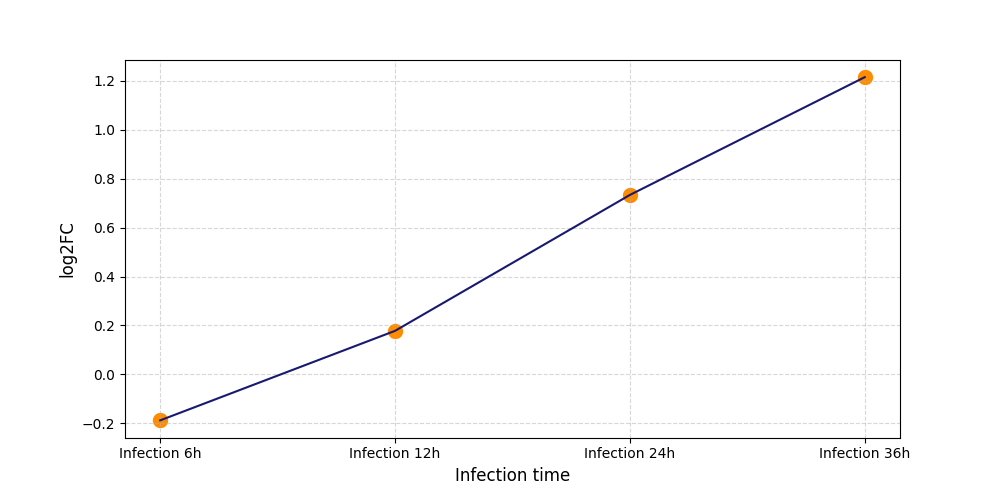 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T603
[19] |
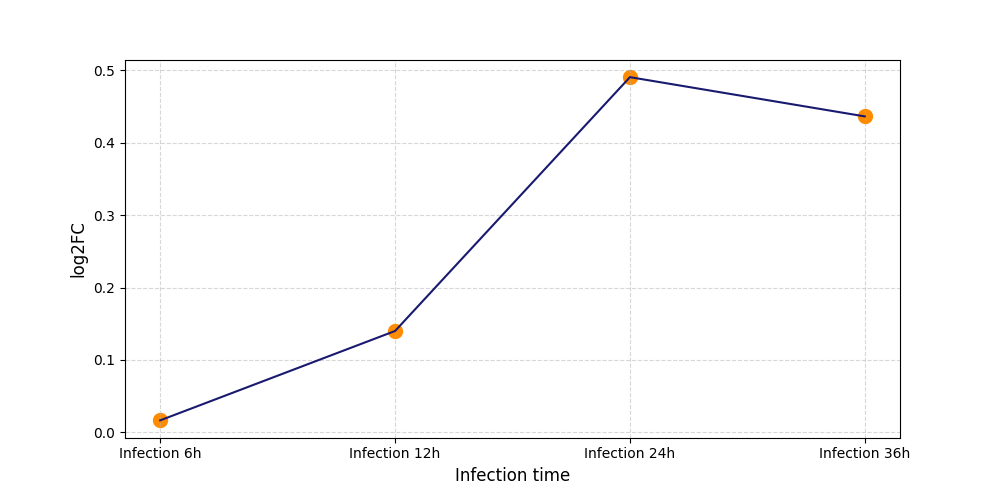 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
T607
[19] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MNPRQGYSLSGYYTHPFQGYEHRQLRYQQPGPGSSPSSFLLKQIEFLKGQLPEAPVIGKQTPSLPPSLPGLRPRFPVLLASSTRGRQVDIRGVPRGVHLRSQGLQRGFQHPSPRGRSLPQRGVDCLSSHFQELSIYQDQEQRILKFLEELGEGKATTAHDLSGKLGTPKKEINRVLYSLAKKGKLQKEAGTPPLWKIAVSTQAWNQHSGVVRPDGHSQGAPNSDPSLEPEDRNSTSVSEDLLEPFIAVSAQAWNQHSGVVRPDSHSQGSPNSDPGLEPEDSNSTSALEDPLEFLDMAEIKEKICDYLFNVSDSSALNLAKNIGLTKARDINAVLIDMERQGDVYRQGTTPPIWHLTDKKRERMQIKRNTNSVPETAPAAIPETKRNAEFLTCNIPTSNASNNMVTTEKVENGQEPVIKLENRQEARPEPARLKPPVHYNGPSKAGYVDFENGQWATDDIPDDLNSIRAAPGEFRAIMEMPSFYSHGLPRCSPYKKLTECQLKNPISGLLEYAQFASQTCEFNMIEQSGPPHEPRFKFQVVINGREFPPAEAGSKKVAKQDAAMKAMTILLEEAKAKDSGKSEESSHYSTEKESEKTAESQTPTPSATSFFSGKSPVTTLLECMHKLGNSCEFRLLSKEGPAHEPKFQYCVAVGAQTFPSVSAPSKKVAKQMAAEEAMKALHGEATNSMASDNQPEGMISESLDNLESMMPNKVRKIGELVRYLNTNPVGGLLEYARSHGFAAEFKLVDQSGPPHEPKFVYQAKVGGRWFPAVCAHSKKQGKQEAADAALRVLIGENEKAERMGFTEVTPVTGASLRRTMLLLSRSPEAQPKTLPLTGSTFHDQIAMLSHRCFNTLTNSFQPSLLGRKILAAIIMKKDSEDMGVVVSLGTGNRCVKGDSLSLKGETVNDCHAEIISRRGFIRFLYSELMKYNSQTAKDSIFEPAKGGEKLQIKKTVSFHLYISTAPCGDGALFDKSCSDRAMESTESRHYPVFENPKQGKLRTKVENGEGTIPVESSDIVPTWDGIRLGERLRTMSCSDKILRWNVLGLQGALLTHFLQPIYLKSVTLGYLFSQGHLTRAICCRVTRDGSAFEDGLRHPFIVNHPKVGRVSIYDSKRQSGKTKETSVNWCLADGYDLEILDGTRGTVDGPRNELSRVSKKNIFLLFKKLCSFRYRRDLLRLSYGEAKKAARDYETAKNYFKKGLKDMGYGNWISKPQEEKNFYLCPV
Click to Show/Hide
|
|---|