Details of Host Protein
| Host Protein General Information (ID: PT0654) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
mRNA decay activator protein ZFP36 (ZFP36)
|
Gene Name |
ZFP36
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
TTP_HUMAN
|
||||||
| Subcellular Location |
Nucleus; P-body
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Zinc-finger RNA-binding protein that destabilizes severalcytoplasmic AU-rich element (ARE) -containing mRNA transcripts bypromoting their poly (A) tail removal or deadenylation, and henceprovide a mechanism for attenuating protein synthesis. Acts as an 3'-untranslated region (UTR) ARE mRNA-binding adapterprotein to communicate signaling events to the mRNA decay machinery. Recruits deadenylase CNOT7 (andprobably the CCR4-NOT complex) via association with CNOT1, and hencepromotes ARE-mediated mRNA deadenylation. Functionsalso by recruiting components of the cytoplasmic RNA decay machinery tothe bound ARE-containing mRNAs. Self regulates by destabilizing itsown mRNA. Binds to 3'-UTR ARE of numerous mRNAs andof its own mRNA. Plays a role in anti-inflammatory responses;suppresses tumor necrosis factor (TNF) -alpha production by stimulatingARE-mediated TNF-alpha mRNA decay and several other inflammatory ARE-containing mRNAs in interferon (IFN) - and/or lipopolysaccharide (LPS) -induced macrophages. Plays also a role in theregulation of dendritic cell maturation at the post-transcriptionallevel, and hence operates as part of a negative feedback loop to limitthe inflammatory response. Promotes ARE-mediated mRNAdecay of hypoxia-inducible factor HIF1A mRNA during the response ofendothelial cells to hypoxia. Positively regulatesearly adipogenesis of preadipocytes by promoting ARE-mediated mRNAdecay of immediate early genes (IEGs). Negativelyregulates hematopoietic/erythroid cell differentiation by promotingARE-mediated mRNA decay of the transcription factor STAT5B mRNA. Plays a role in maintaining skeletal musclesatellite cell quiescence by promoting ARE-mediated mRNA decay of themyogenic determination factor MYOD1 mRNA. Associatesalso with and regulates the expression of non-ARE-containing targetmRNAs at the post-transcriptional level, such as MHC class I mRNAs. Participates in association with argonaute RISCcatalytic components in the ARE-mediated mRNA decay mechanism; assistsmicroRNA (miRNA) targeting ARE-containing mRNAs. Mayalso play a role in the regulation of cytoplasmic mRNA decapping;enhances decapping of ARE-containing RNAs, in vitro. Involved in the delivery of target ARE-mRNAs to processing bodies (PBs). In addition to its cytosolic mRNA-decay function, affects nuclear pre-mRNA processing. Negativelyregulates nuclear poly (A) -binding protein PABPN1-stimulatedpolyadenylation activity on ARE-containing pre-mRNA during LPS-stimulated macrophages. Also involved in the regulationof stress granule (SG) and P-body (PB) formation and fusion. Plays a role in the regulation of keratinocyteproliferation, differentiation and apoptosis. Plays arole as a tumor suppressor by inhibiting cell proliferation in breastcancer cells.
[1-5]
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| Human T-cell leukemia virus 1 infection | hsa05166 |
Pathway Map 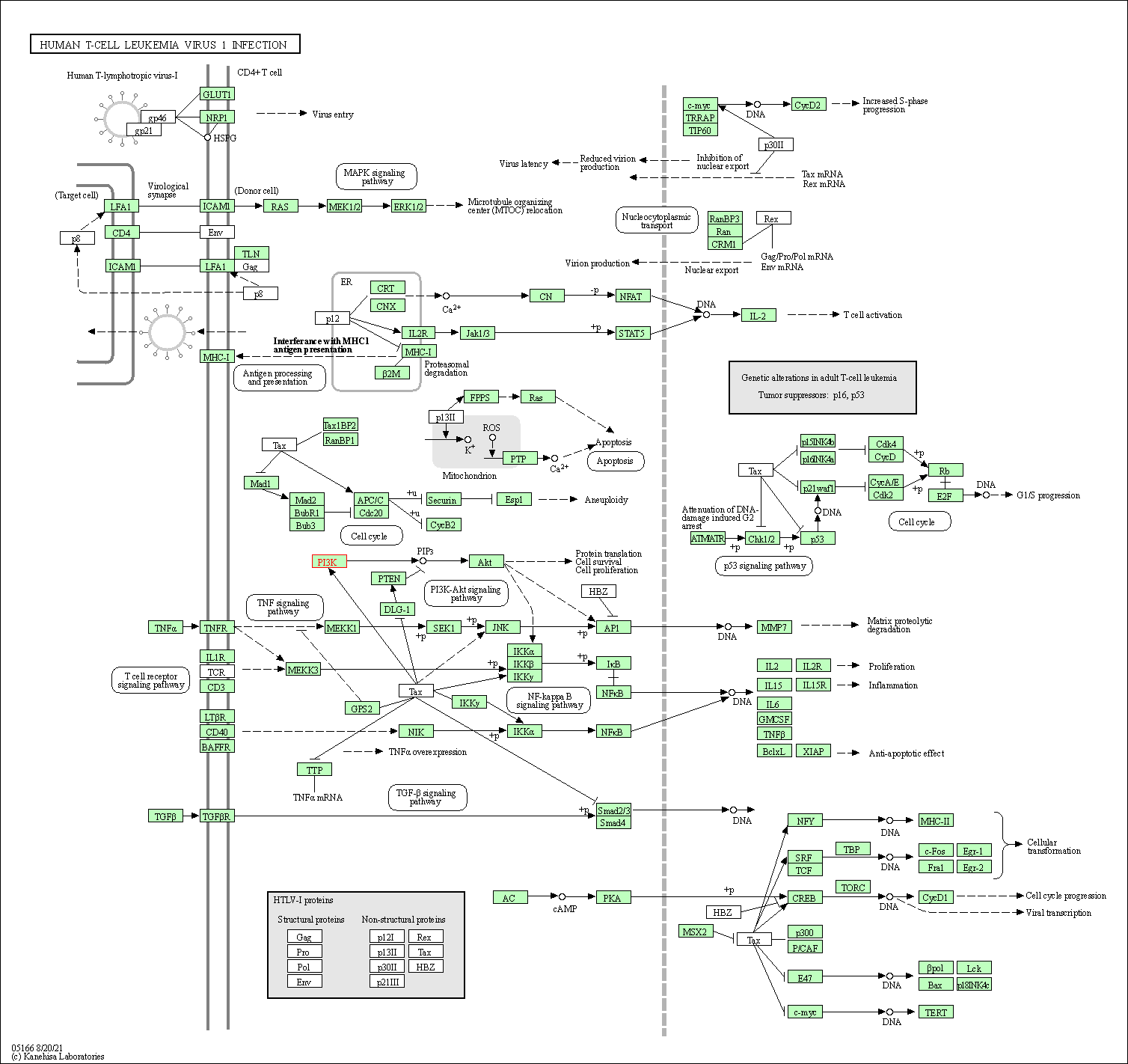
|
|||||||
| Kaposi sarcoma-associated herpesvirus infection | hsa05167 |
Pathway Map 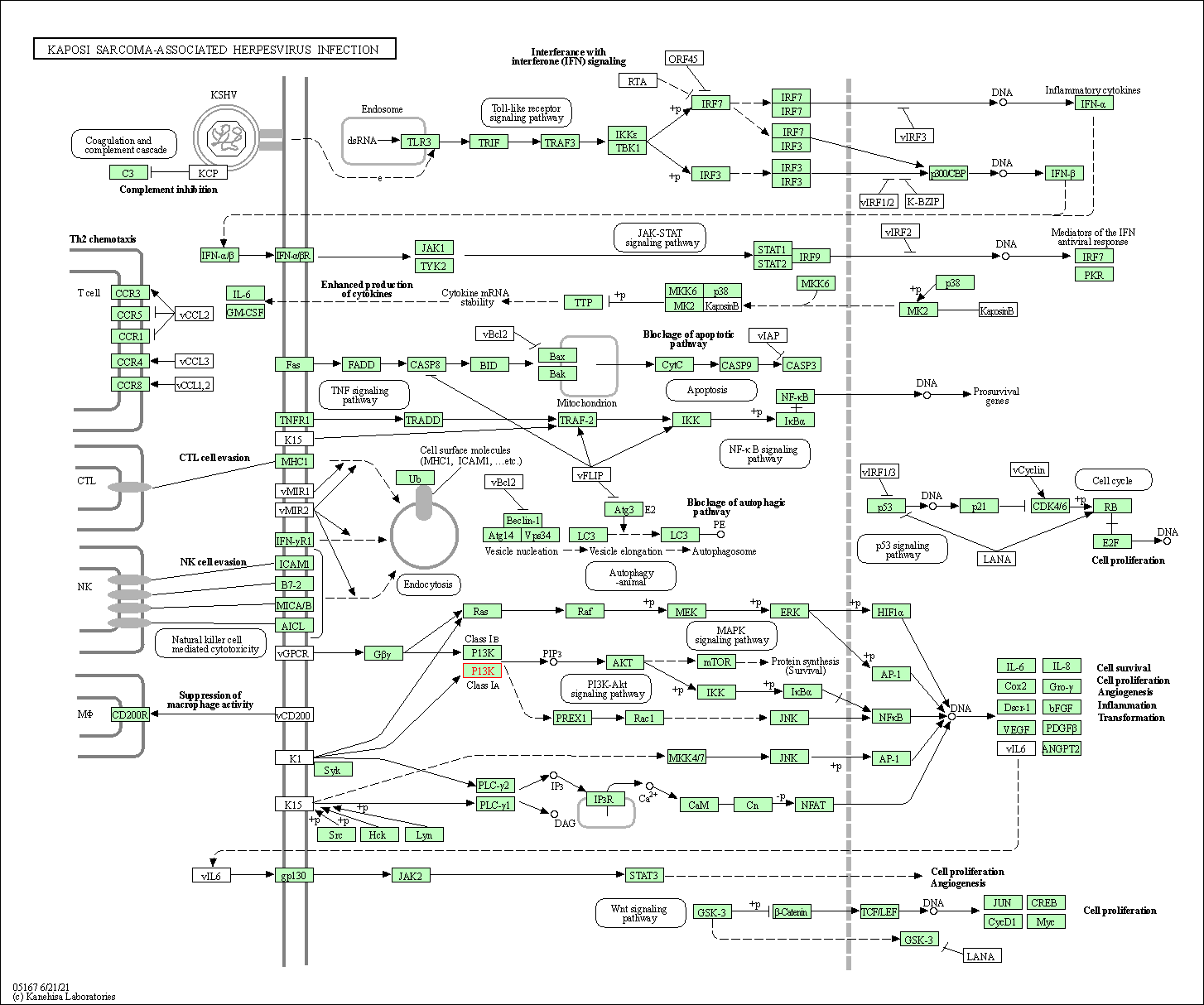
|
|||||||
| 3D Structure |
|
||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: 3'-UTR (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: 5'-UTR (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF10 (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF10
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF1ab (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF3a (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF3a
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF6 (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF6
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF7a (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF7a
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF7b (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF7b
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF8 (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF8
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: E region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: M region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: N region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: S region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/England/02/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[7] | |||||||
| Strains Name |
hCoV-19/England/02/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Calu-3 cells (Human lung cancer cell) Calu-3 cells (Human lung cancer cell) (CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Lung | ||||||||
| Infection Time | 24 h | ||||||||
| Interaction Score | P-adjust = 0.008 | ||||||||
| Method Description | UV protein-RNA crosslinking; RNA interactome capture (cRIC); RNA antisense purification coupled with mass spectrometry (RAP-MS) | ||||||||
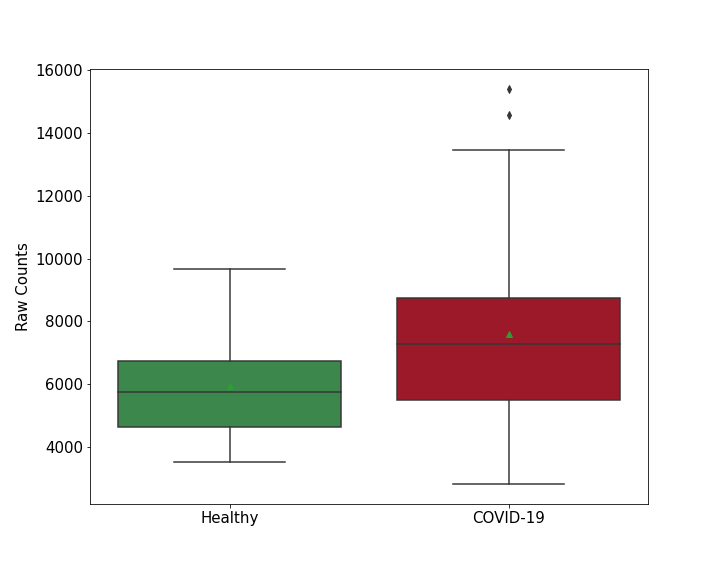
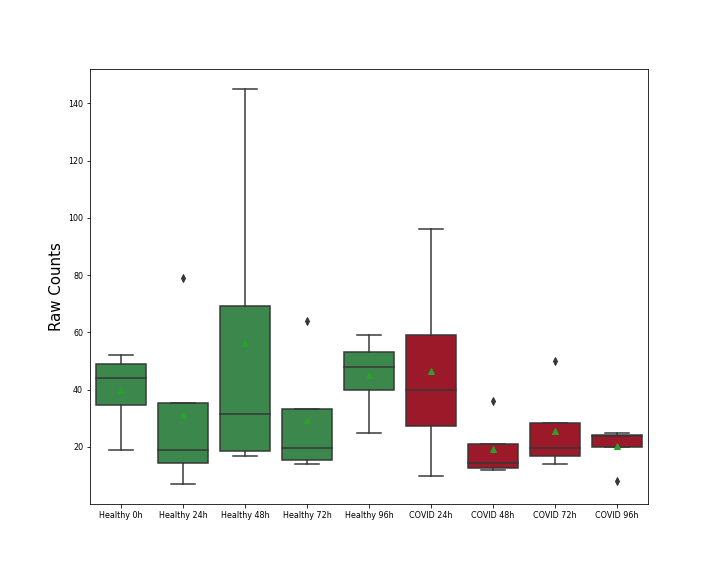
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
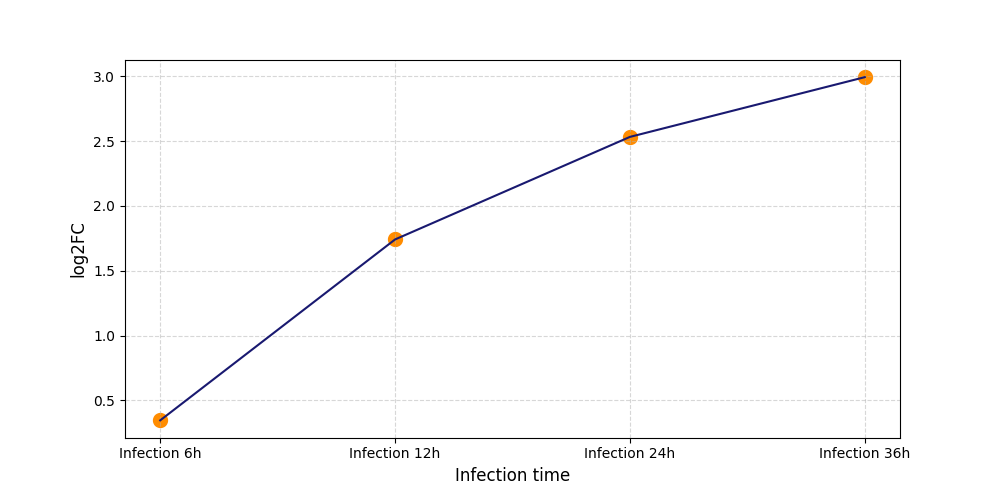
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S186
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MDLTAIYESLLSLSPDVPVPSDHGGTESSPGWGSSGPWSLSPSDSSPSGVTSRLPGRSTSLVEGRSCGWVPPPPGFAPLAPRLGPELSPSPTSPTATSTTPSRYKTELCRTFSESGRCRYGAKCQFAHGLGELRQANRHPKYKTELCHKFYLQGRCPYGSRCHFIHNPSEDLAAPGHPPVLRQSISFSGLPSGRRTSPPPPGLAGPSLSSSSFSPSSSPPPPGDLPLSPSAFSAAPGTPLARRDPTPVCCPSCRRATPISVWGPLGGLVRTPSVQSLGSDPDEYASSGSSLGGSDSPVFEAGVFAPPQPVAAPRRLPIFNRISVSE
Click to Show/Hide
|
|---|