Details of Host Protein
| Host Protein General Information (ID: PT1263) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
YTH domain-containing protein 1 (YTHDC1)
|
Gene Name |
YTHDC1
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
YTDC1_HUMAN
|
||||||
| Subcellular Location |
Nucleus speckle
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Regulator of alternative splicing that specificallyrecognizes and binds N6-methyladenosine (m6A)-containing RNAs. M6A is a modification present at internal sites of mRNAs and some non-coding RNAs and plays a role in the efficiency of mRNA splicing, processing and stability. Acts as akey regulator of exon-inclusion or exon-skipping during alternativesplicing via interaction with mRNA splicing factors SRSF3 and SRSF10. Specifically binds m6A-containing mRNAs and promotesrecruitment of SRSF3 to its mRNA-binding elements adjacent to m6Asites, leading to exon-inclusion during alternative splicing. In contrast, interaction with SRSF3 preventsinteraction with SRSF10, a splicing factor that promotes exon skipping:this prevents SRSF10 from binding to its mRNA-binding sites close tom6A-containing regions, leading to inhibit exon skipping duringalternative splicing. May also regulate alternativesplice site selection. Also involved in nuclearexport of m6A-containing mRNAs via interaction with SRSF3: interactionwith SRSF3 facilitates m6A-containing mRNA-binding to both SRSF3 andNXF1, promoting mRNA nuclear export. Involved in S-adenosyl-L-methionine homeostasis by regulating expression of MAT2Atranscripts, probably by binding m6A-containing MAT2A mRNAs. Also recognizes and binds m6A on other RNA molecules. Involved in random X inactivation mediated by XistRNA: recognizes and binds m6A-containing Xist and promotestranscription repression activity of Xist. Alsorecognizes and binds m6A-containing single-stranded DNA. Involved in germline development: required forspermatogonial development in males and oocyte growth and maturation infemales, probably via its role in alternative splicing.
[1-4]
Click to Show/Hide
|
||||||||
| 3D Structure |
|
||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: 3'-UTR (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: 5'-UTR (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF10 (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF10
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF1ab (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF3a (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF3a
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF6 (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF6
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF7a (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF7a
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF8 (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF8
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: E region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: M region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: N region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: S region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/England/02/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/England/02/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Calu-3 cells (Human lung cancer cell) Calu-3 cells (Human lung cancer cell) (CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Lung | ||||||||
| Infection Time | 24 h | ||||||||
| Interaction Score | P-adjust = 0.008 | ||||||||
| Method Description | UV protein-RNA crosslinking; RNA interactome capture (cRIC); RNA antisense purification coupled with mass spectrometry (RAP-MS) | ||||||||
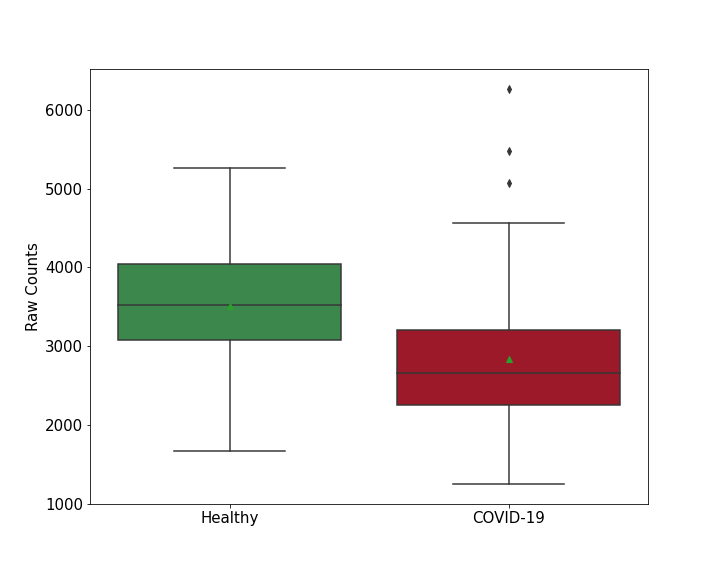
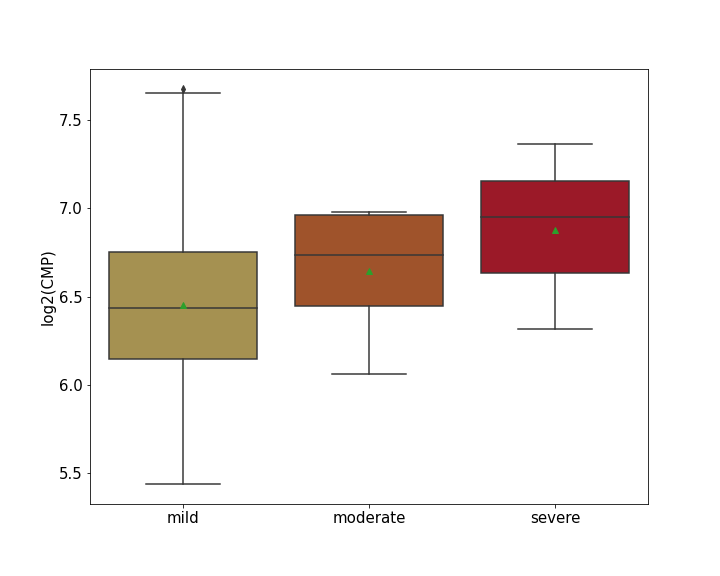
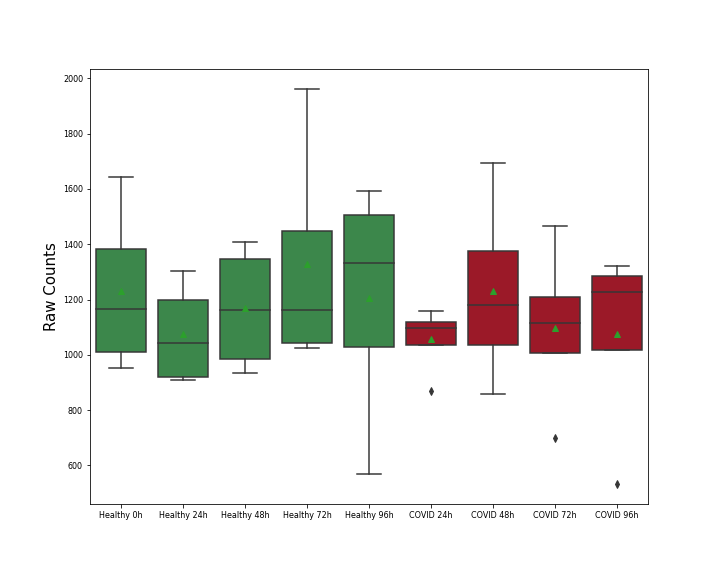
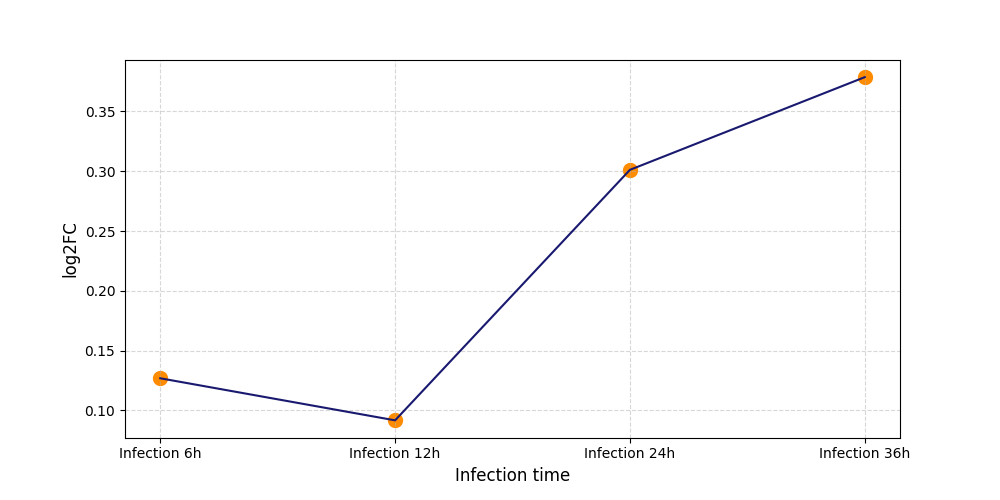
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
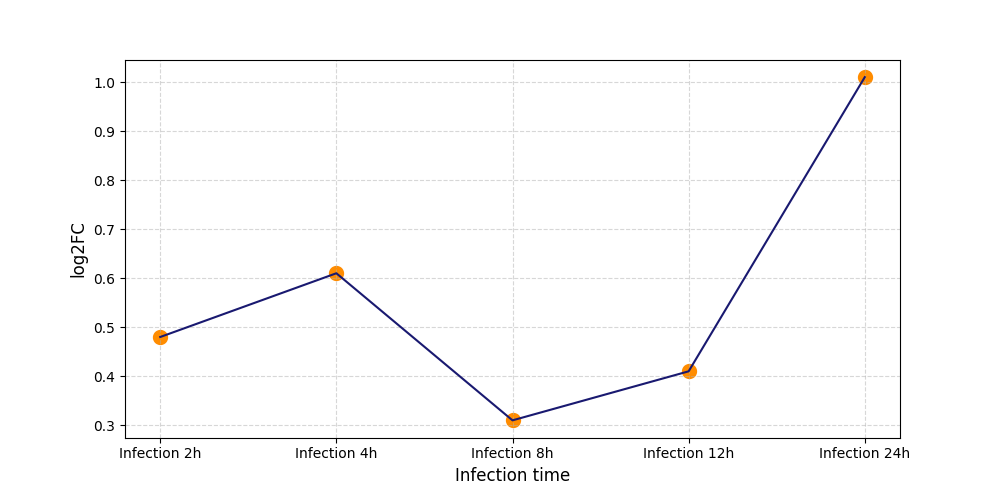
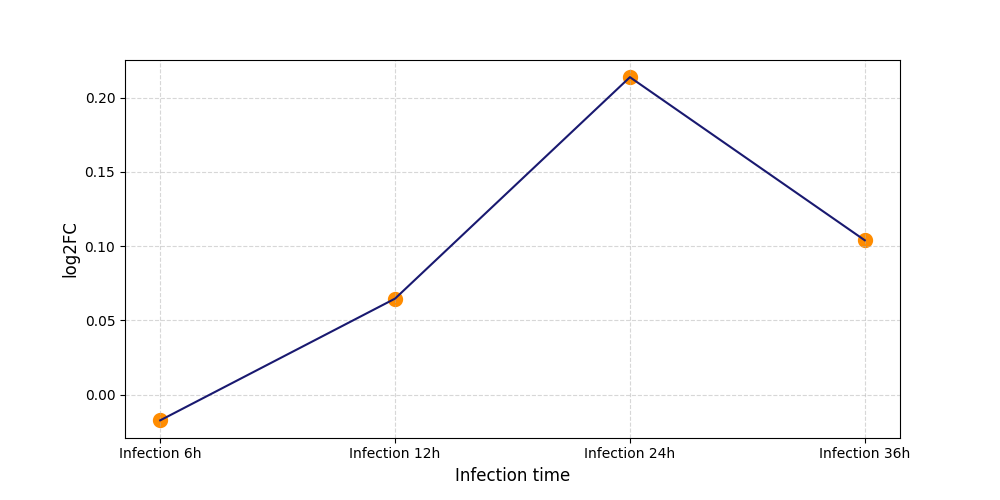
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S146
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
S146
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
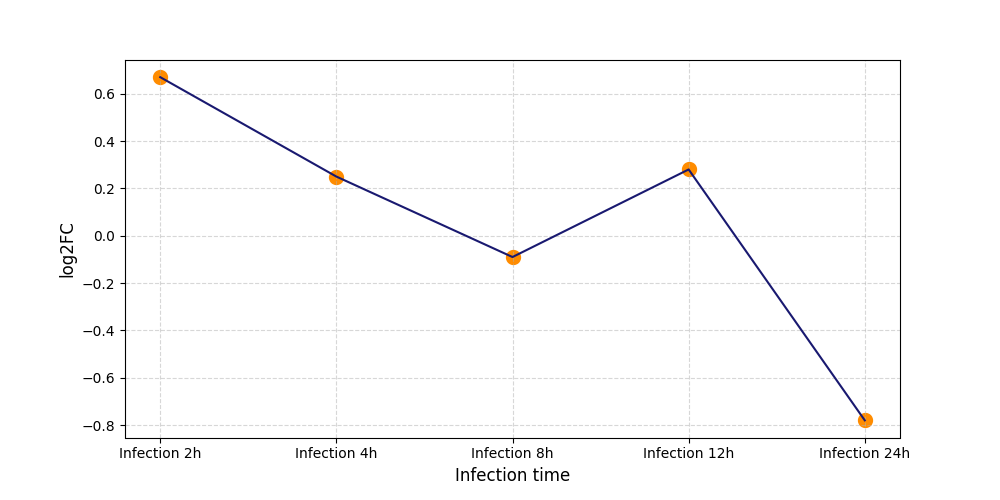
S308
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
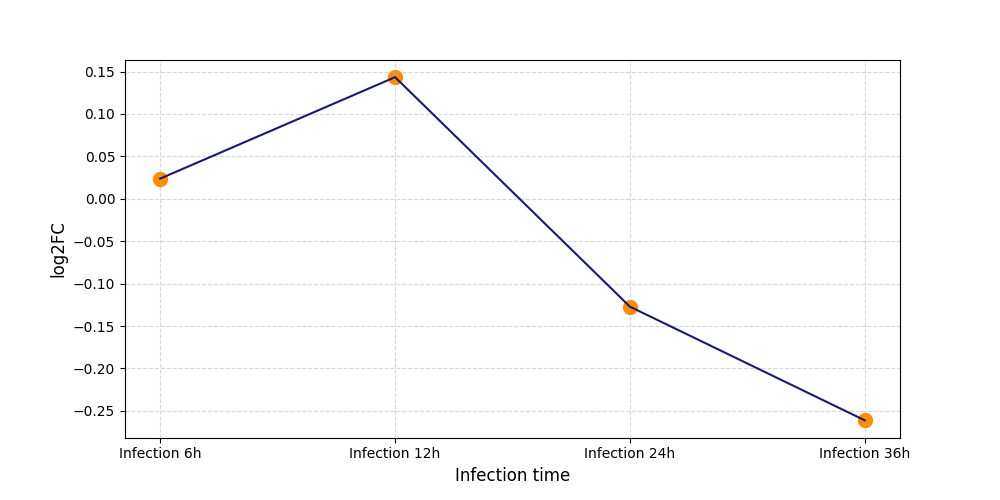
S308
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
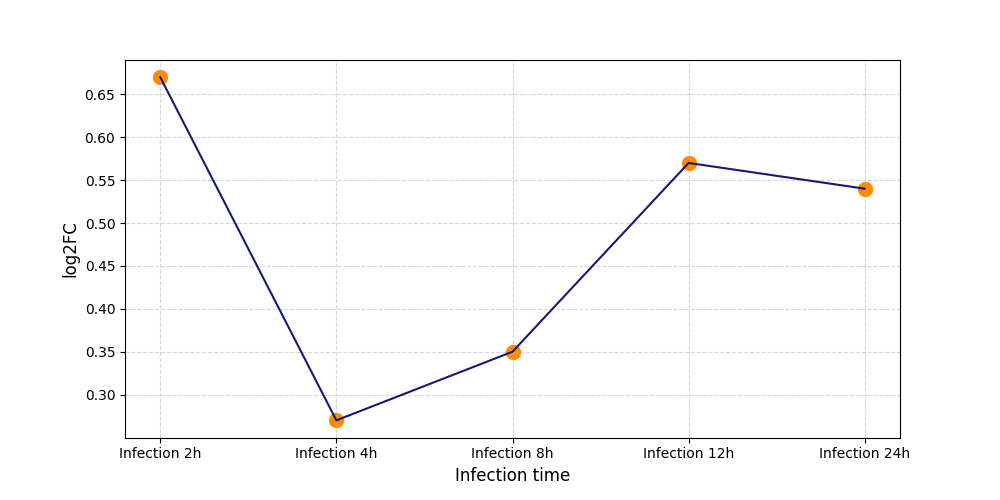
S424
[7] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
S424
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MAADSREEKDGELNVLDDILTEVPEQDDELYNPESEQDKNEKKGSKRKSDRMESTDTKRQKPSVHSRQLVSKPLSSSVSNNKRIVSTKGKSATEYKNEEYQRSERNKRLDADRKIRLSSSASREPYKNQPEKTCVRKRDPERRAKSPTPDGSERIGLEVDRRASRSSQSSKEEVNSEEYGSDHETGSSGSSDEQGNNTENEEEGVEEDVEEDEEVEEDAEEDEEVDEDGEEEEEEEEEEEEEEEEEEEEYEQDERDQKEEGNDYDTRSEASDSGSESVSFTDGSVRSGSGTDGSDEKKKERKRARGISPIVFDRSGSSASESYAGSEKKHEKLSSSVRAVRKDQTSKLKYVLQDARFFLIKSNNHENVSLAKAKGVWSTLPVNEKKLNLAFRSARSVILIFSVRESGKFQGFARLSSESHHGGSPIHWVLPAGMSAKMLGGVFKIDWICRRELPFTKSAHLTNPWNEHKPVKIGRDGQEIELECGTQLCLLFPPDESIDLYQVIHKMRHKRRMHSQPRSRGRPSRREPVRDVGRRRPEDYDIHNSRKKPRIDYPPEFHQRPGYLKDPRYQEVDRRFSGVRRDVFLNGSYNDYVREFHNMGPPPPWQGMPPYPGMEQPPHHPYYQHHAPPPQAHPPYSGHHPVPHEARYRDKRVHDYDMRVDDFLRRTQAVVSGRRSRPRERDRERERDRPRDNRRDRERDRGRDRERERERLCDRDRDRGERGRYRR
Click to Show/Hide
|
|---|