Details of Host Protein
| Host Protein General Information (ID: PT1271) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
Zinc finger CCCH domain-containing protein 8 (ZC3H8)
|
Gene Name |
ZC3H8
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
ZC3H8_HUMAN
|
||||||
| Subcellular Location |
Nucleus
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Acts as a transcriptional repressor of the GATA3 promoter. Sequence-specific DNA-binding factor that binds to the 5'-AGGTCTC-3'sequence within the negative cis-acting element intronic regulatoryregion (IRR) of the GATA3 gene. Component of the littleelongation complex (LEC), a complex required to regulate small nuclearRNA (snRNA) gene transcription by RNA polymerase II and III. Induces thymocyte apoptosis when overexpressed, which may indicate a role in regulation of thymocyte homeostasis.
[1-3]
Click to Show/Hide
|
||||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Anti-viral | [4] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein ZC3H8 in viral infection, ZC3H8 protein Knockout Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of ZC3H8 increases SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: ORF10 (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
ORF10
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (human liver cell line) (CVCL_0336 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| RNA Region: ORF1ab (hCoV-19/Venezuela/Bol285/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Venezuela/Bol285/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF1ab (hCoV-19/Switzerland/TI-EOC-902_25839324/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Switzerland/TI-EOC-902_25839324/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF1ab (hCoV-19/Suriname/SR-342/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Suriname/SR-342/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF1ab (hCoV-19/Peru/CAL-INS-5458/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Peru/CAL-INS-5458/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF1ab (hCoV-19/Bolivia/CH-INLASA-27102/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Bolivia/CH-INLASA-27102/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF1ab (hCoV-19/Argentina/PAIS-E0546/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Argentina/PAIS-E0546/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF1ab (hCoV-19/Argentina/INEI113521/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Argentina/INEI113521/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF1ab (hCoV-19/Argentina/INEI106123/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Argentina/INEI106123/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF1ab (hCoV-19/Argentina/INEI104133/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Argentina/INEI104133/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF8 (hCoV-19/Bolivia/CH-INLASA-27102/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Bolivia/CH-INLASA-27102/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF8
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF8 (hCoV-19/Argentina/INEI106123/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Argentina/INEI106123/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF8
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-SARS/Manitoba/2003 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-SARS/Manitoba/2003
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome-related coronavirus (HCoV-SARS)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-OC43/VR-759 Quebec/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-OC43/VR-759 Quebec/2019
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus OC43 (HCov-OC43)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-NL63/Amsterdam/2004 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-NL63/Amsterdam/2004
|
||||||||
| Strains Family |
Alpha
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human Coronavirus NL63 (HCov-NL63)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-MERS/Jeddah/2012 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-MERS/Jeddah/2012
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Middle East respiratory syndrome-related coronavirus (HCov-MERS)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-HKU1/Hong Kong/2004 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-HKU1/Hong Kong/2004
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus HKU1 (HCoV-HKU1)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-229E/Würzburg/2000 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-229E/Würzburg/2000
|
||||||||
| Strains Family |
Alpha
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus 229E (HCov-229E)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | K562 cells (Human leukemic cell line) K562 cells (Human leukemic cell line) (CVCL_0004 ) | ||||||||
| Cell Originated Tissue | Bone marrow | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/England/02/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[7] | |||||||
| Strains Name |
hCoV-19/England/02/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Calu-3 cells (Human lung cancer cell) Calu-3 cells (Human lung cancer cell) (CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Lung | ||||||||
| Infection Time | 24 h | ||||||||
| Interaction Score | P-adjust = 0.022 | ||||||||
| Method Description | UV protein-RNA crosslinking; RNA interactome capture (cRIC); RNA antisense purification coupled with mass spectrometry (RAP-MS) | ||||||||
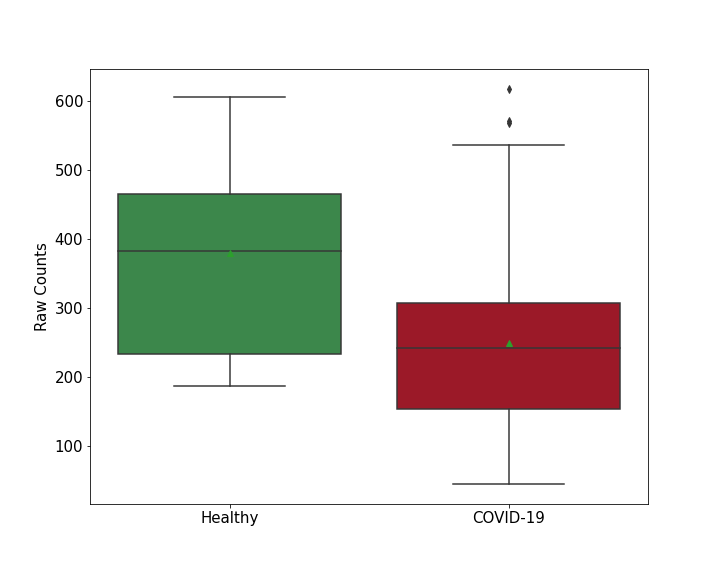
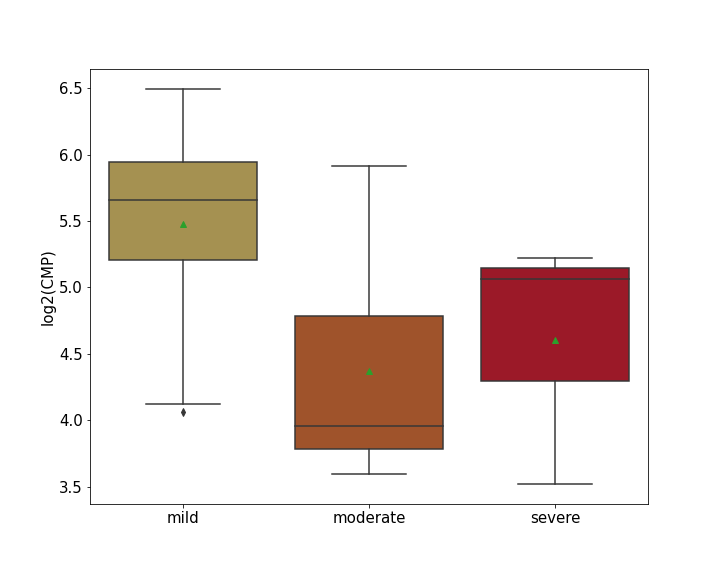
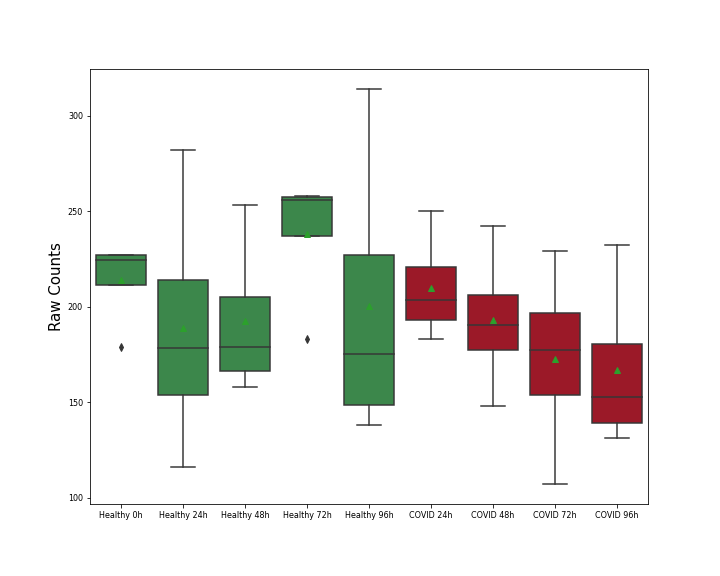
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
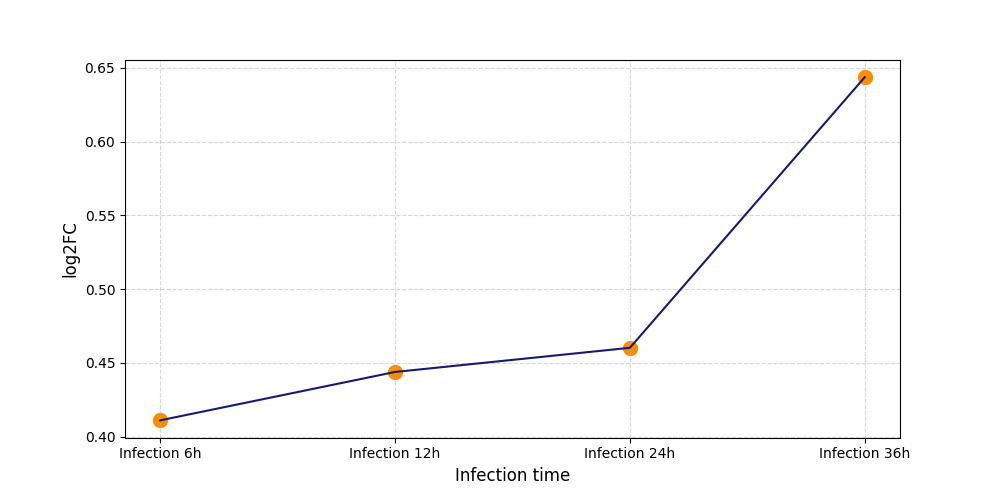
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S59
[8] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MDFENLFSKPPNPALGKTATDSDERIDDEIDTEVEETQEEKIKLECEQIPKKFRHSAISPKSSLHRKSRSKDYDVYSDNDICSQESEDNFAKELQQYIQAREMANAAQPEESTKKEGVKDTPQAAKQKNKNLKAGHKNGKQKKMKRKWPGPGNKGSNALLRNSGSQEEDGKPKEKQQHLSQAFINQHTVERKGKQICKYFLERKCIKGDQCKFDHDAEIEKKKEMCKFYVQGYCTRGENCLYLHNEYPCKFYHTGTKCYQGEYCKFSHAPLTPETQELLAKVLDTEKKSCK
Click to Show/Hide
|
|---|