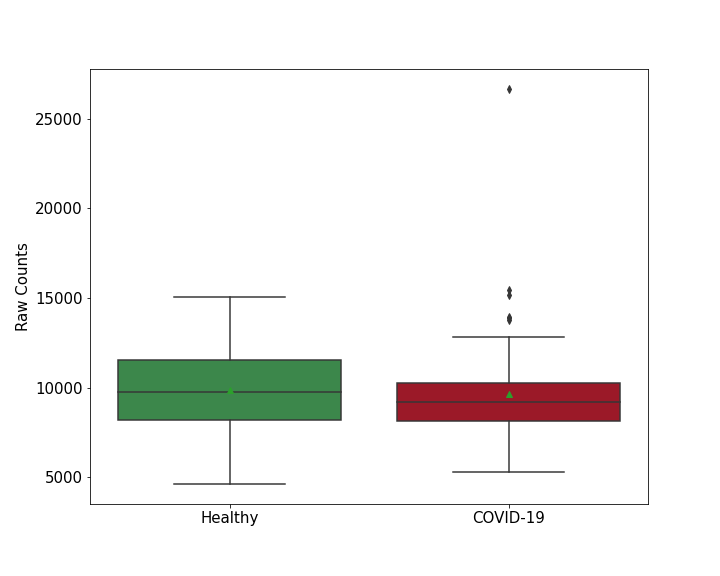
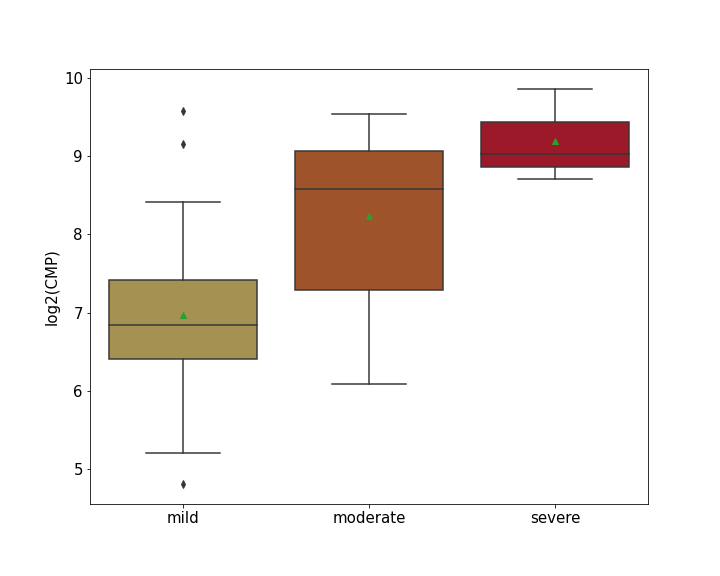
Details of Host Protein
| Host Protein General Information (ID: PT0210) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
Helicase-like protein 2 (HLP2)
|
Gene Name |
DDX3X
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
DDX3X_HUMAN
|
||||||
| Protein Families |
DEAD box helicase family
|
||||||||
| EC Number |
3.6.4.13
|
||||||||
| Subcellular Location |
Nucleus; Cytoplasm
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Multifunctional ATP-dependent RNA helicase. The ATPase activity can bestimulated by various ribo-and deoxynucleic acids indicative for arelaxed substrate specificity. In vitro can unwindpartially double-stranded DNA with a preference for 5'-single-strandedDNA overhangs. Binds RNA G-quadruplex (rG4s) structures, including those located in the 5'-UTR ofNRAS mRNA. Involved in many cellular processes, whichdo not necessarily require its ATPase/helicase catalytic activities. Involved in transcription regulation. Positively regulates CDKN1A/WAF1/CIP1 transcriptionin an SP1-dependent manner, hence inhibits cell growth. This functionrequires its ATPase, but not helicase activity. CDKN1A up-regulation may be cell-type specific. Binds CDH1/E-cadherin promoter and represses itstranscription. Potentiates HNF4A-mediated MTTPtranscriptional activation; this function requires ATPase, but nothelicase activity. Facilitates HNF4A acetylation, possibly catalyzed byCREBBP/EP300, thereby increasing the DNA-binding affinity of HNF4 toits response element. In addition, disrupts the interaction betweenHNF4 and SHP that forms inactive heterodimers and enhances theformation of active HNF4 homodimers. By promoting HNF4A-induced MTTPexpression, may play a role in lipid homeostasis. Maypositively regulate TP53 transcription. Associateswith mRNPs, predominantly with spliced mRNAs carrying an exon junctioncomplex (EJC). Involved in theregulation of translation initiation. Not involved in the general process of translation, but promotes efficient translation of selected complex mRNAs, containing highly structured 5'-untranslated regions (UTR). This function depends on helicaseactivity. Might facilitatetranslation by resolving secondary structures of 5'-UTRs duringribosome scanning. Alternatively, may act prior to43S ribosomal scanning and promote 43S pre-initiation complex entry tomRNAs exhibiting specific RNA motifs, by performing local remodeling oftranscript structures located close to the cap moiety. Independently of its ATPase activity, promotes theassembly of functional 80S ribosomes and disassembles from ribosomesprior to the translation elongation process. Positively regulates the translation of cyclin E1/CCNE1 mRNA andconsequently promotes G1/S-phase transition during the cell cycle. May activate TP53 translation. Required for endoplasmic reticulum stress-induced ATF4 mRNA translation. Independently of its ATPase/helicase activity, enhances IRES-mediated translation; this activity requires interactionwith EIF4E. Independently of itsATPase/helicase activity, has also been shown specifically repress cap-dependent translation, possibly by acting on translation initiationfactor EIF4E. Involved in innate immunity, acting asa viral RNA sensor. Binds viral RNAs and promotes the production oftype I interferon (IFN-alpha and IFN-beta). Potentiate MAVS/DDX58-mediatedinduction of IFNB in early stages of infection. Enhances IFNB1 expression via IRF3/IRF7 pathway andparticipates in NFKB activation in the presence of MAVS and TBK1. Involved in TBK1 and IKBKE-dependent IRF3 activationleading to IFNB induction, acts as a scaffolding adapter that linksIKBKE and IRF3 and coordinates their activation. Involved in the TLR7/TLR8 signaling pathway leading to type Iinterferon induction, including IFNA4 production. In this context, actsas an upstream regulator of IRF7 activation by MAP3K14/NIK andCHUK/IKKA. Stimulates CHUK autophosphorylation and activation followingphysiological activation of the TLR7 and TLR8 pathways, leading toMAP3K14/CHUK-mediated activatory phosphorylation of IRF7. Also stimulates MAP3K14/CHUK-dependent NF-kappa-Bsignaling. Negatively regulates TNF-induced IL6 andIL8 expression, via the NF-kappa-B pathway. May act by interacting withRELA/p65 and trapping it in the cytoplasm. May alsobind IFNB promoter; the function is independent of IRF3. Involved in both stress and inflammatory responses. Independently of its ATPase/helicase activity, required for efficient stress granule assembly through its interactionwith EIF4E, hence promotes survival in stressed cells. Independently of its helicase activity, regulatesNLRP3 inflammasome assembly through interaction with NLRP3 and hencepromotes cell death by pyroptosis during inflammation. This function isindependent of helicase activity. Therefore DDX3Xavailability may be used to interpret stress signals and choose betweenpro-survival stress granules and pyroptotic NLRP3 inflammasomes andserve as a live-or-die checkpoint in stressed cells. Inassociation with GSK3A/B, negatively regulates extrinsic apoptoticsignaling pathway via death domain receptors, including TNFRSF10B, slowing down the rate of CASP3 activation following death receptorstimulation. Cleavage by caspases may inactivateDDX3X and relieve the inhibition. Independently ofits ATPase/helicase activity, allosteric activator of CSNK1E. Stimulates CSNK1E-mediated phosphorylation of DVL2, thereby involved inthe positive regulation of Wnt/beta-catenin signaling pathway. Alsoactivates CSNK1A1 and CSNK1D in vitro, but it is uncertain if thesetargets are physiologically relevant. ATPase and casein kinase-activating functions aremutually exclusive. May be involved in mitoticchromosome segregation.
[1-4]
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| RIG-I-like receptor signaling pathway | hsa04622 |
Pathway Map 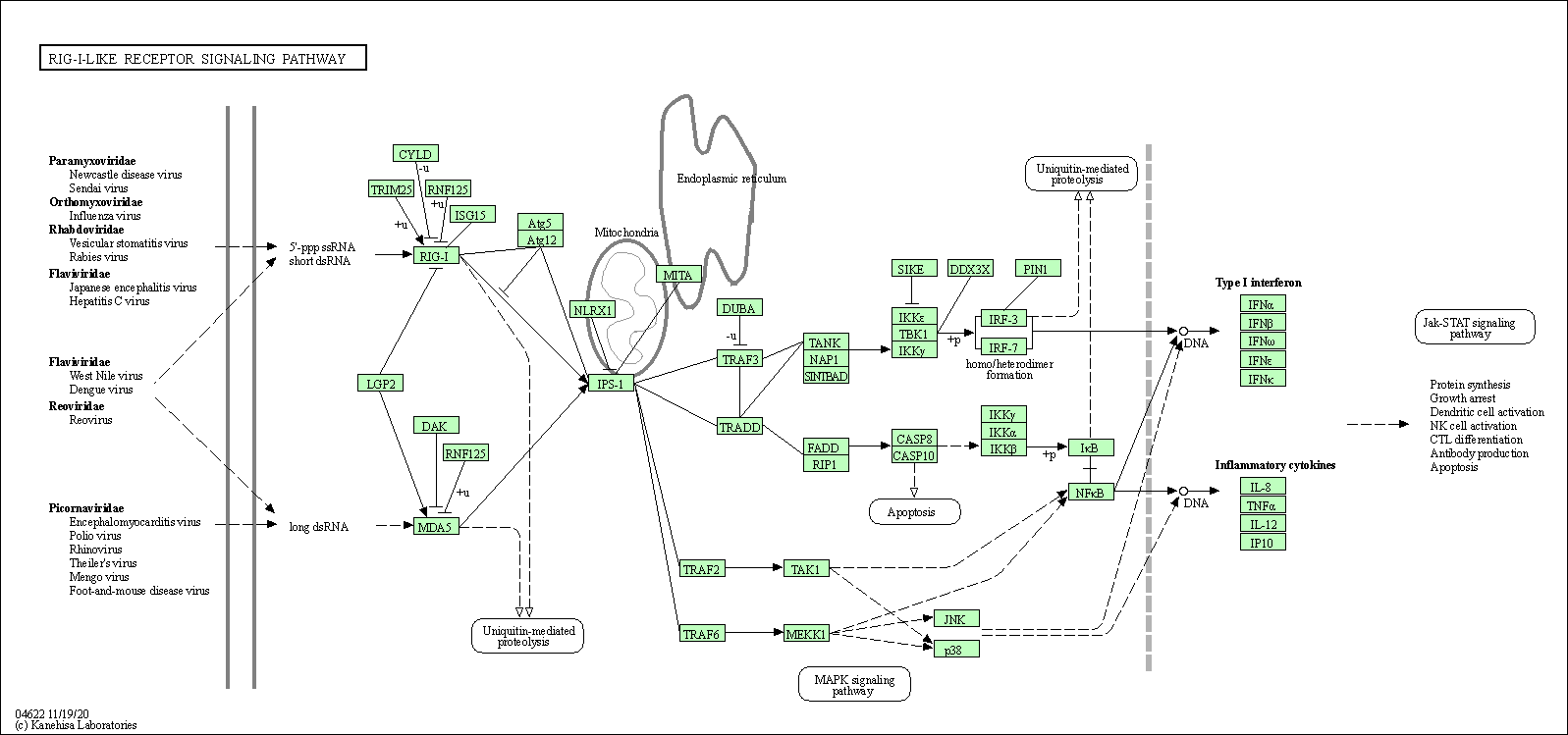
|
|||||||
| Hepatitis B | hsa05161 |
Pathway Map 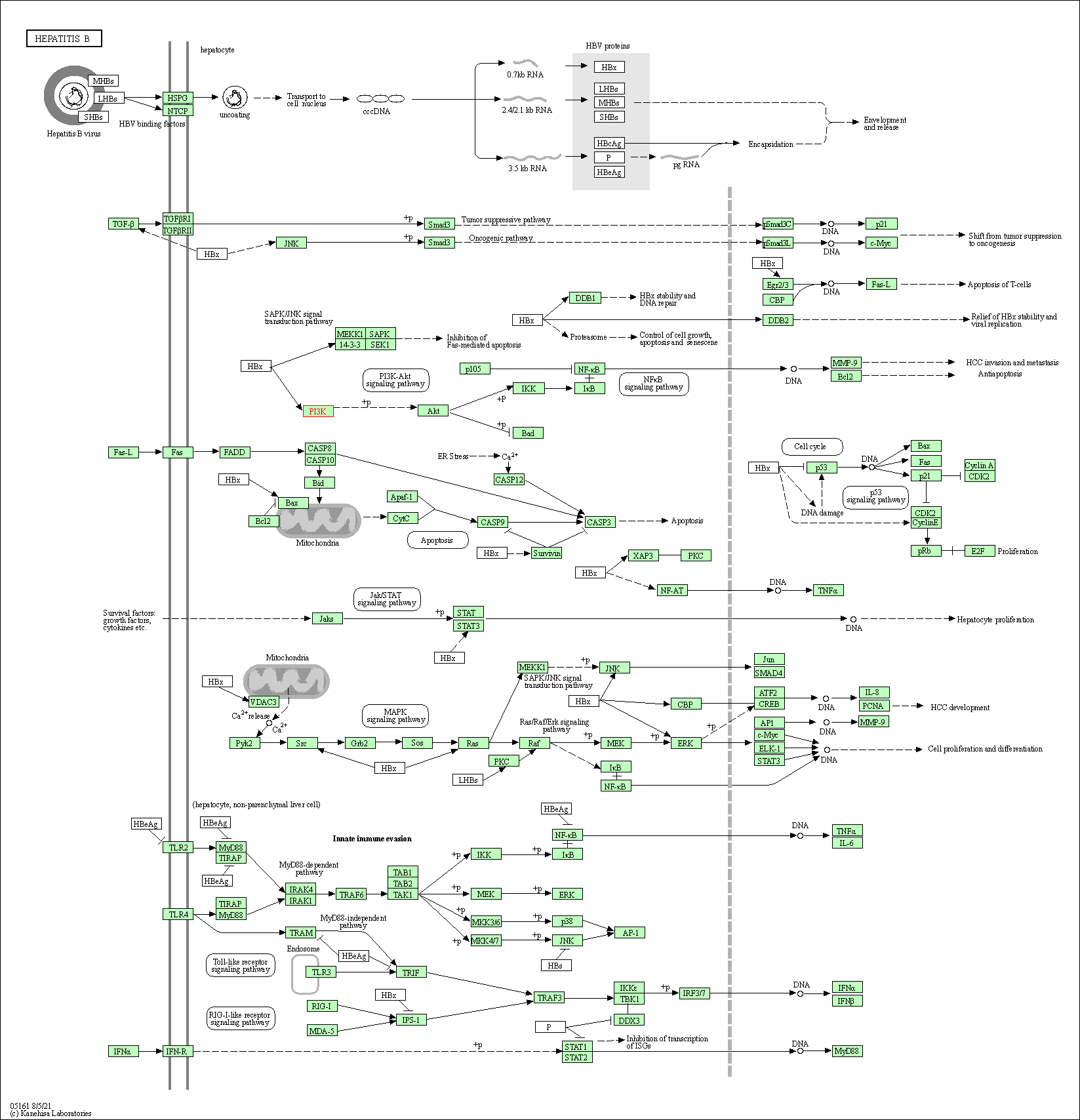
|
|||||||
| Viral carcinogenesis | hsa05203 |
Pathway Map 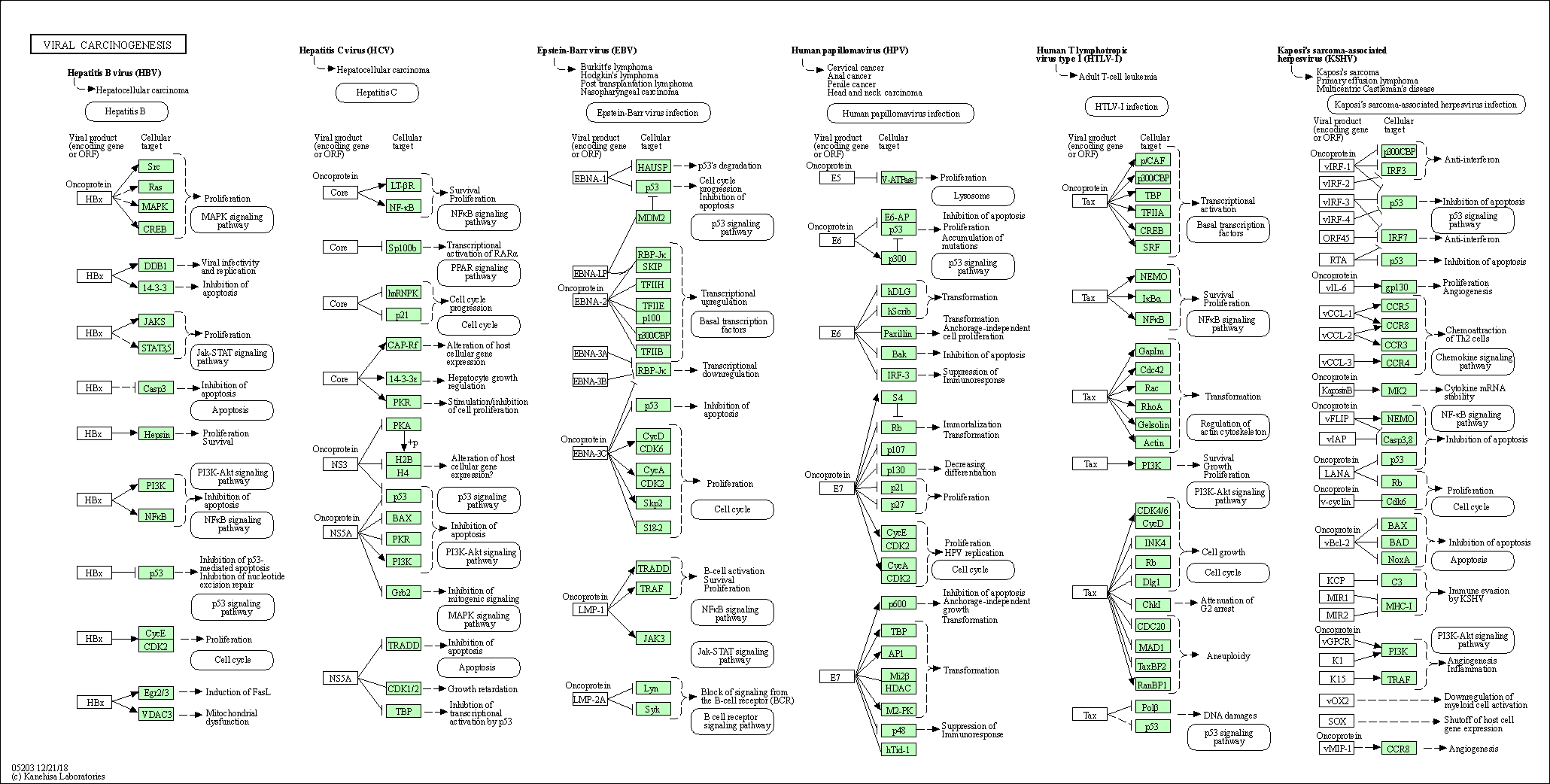
|
|||||||
| 3D Structure |
|
||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: 3'-UTR (hCoV-SARS/Manitoba/2003 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-SARS/Manitoba/2003
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome-related coronavirus (HCoV-SARS)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 3'-UTR (hCov-OC43/VR-759 Quebec/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCov-OC43/VR-759 Quebec/2019
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Human coronavirus OC43 (HCov-OC43)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 3'-UTR (hCov-NL63/Amsterdam/2004 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCov-NL63/Amsterdam/2004
|
||||||||
| Strains Family |
Alpha
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Human Coronavirus NL63 (HCov-NL63)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 3'-UTR (hCov-MERS/Jeddah/2012 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCov-MERS/Jeddah/2012
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Middle East respiratory syndrome-related coronavirus (HCov-MERS)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 3'-UTR (hCoV-HKU1/Hong Kong/2004 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-HKU1/Hong Kong/2004
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Human coronavirus HKU1 (HCoV-HKU1)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 3'-UTR (hCov-229E/Würzburg/2000 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCov-229E/Würzburg/2000
|
||||||||
| Strains Family |
Alpha
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Human coronavirus 229E (HCov-229E)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 3'-UTR (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 3'-UTR (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) (CVCL_0336;CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Liver; Lung | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| RNA Region: 5'-UTR (hCoV-19/South Korea/KCDC03/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[7] | |||||||
| Strains Name |
hCoV-19/South Korea/KCDC03/2020
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Directly bind to SARS-CoV-23 RNA's 5' UTR region | ||||||||
| Infection Cells | Vero cells (epithelial kidney cell) Vero cells (epithelial kidney cell) (CVCL_0059 ) | ||||||||
| Cell Originated Tissue | kidney | ||||||||
| Infection Time | 24 h | ||||||||
| Interaction Score | P-adjust < 0.05 | ||||||||
| Method Description | Modified RNA antisense purification coupled with mass spectrometry (RAP-MS); label-free quantification (LFQ); liquid chromatography with tandem mass spectrometry (LC-MS/MS) | ||||||||
| RNA Region: 5'-UTR (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) (CVCL_0336;CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Liver; Lung | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| RNA Region: 5'-UTR (hCoV-19/Namibia/N17380/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Namibia/N17380/2021
|
||||||||
| Strains Family |
Delta (B.1.617.2 and AY lineages)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 5'-UTR (hCoV-19/Namibia/CERI-KRISP-K022530/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Namibia/CERI-KRISP-K022530/2021
|
||||||||
| Strains Family |
Delta (B.1.617.2 and AY lineages)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 5'-UTR (hCoV-19/Kyrgyzstan/ChVir26576_10/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Kyrgyzstan/ChVir26576_10/2021
|
||||||||
| Strains Family |
Delta (B.1.617.2 and AY lineages)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 5'-UTR (hCoV-19/Kazakhstan/1872/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Kazakhstan/1872/2021
|
||||||||
| Strains Family |
Delta (B.1.617.2 and AY lineages)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 5'-UTR (hCoV-19/Iraq/Thi-Qar-1/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Iraq/Thi-Qar-1/2021
|
||||||||
| Strains Family |
Delta (B.1.617.2 and AY lineages)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF7a (hCoV-19/Kazakhstan/1872/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Kazakhstan/1872/2021
|
||||||||
| Strains Family |
Delta (B.1.617.2 and AY lineages)
|
||||||||
| RNA Binding Region |
ORF7a
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: ORF8 (hCoV-19/Argentina/INEI113521/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Argentina/INEI113521/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
ORF8
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Zambia/ZMB-88673/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Zambia/ZMB-88673/2021
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Wuhan/WHUH020/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan/WHUH020/2020
|
||||||||
| Strains Family |
Epsilon (B.1.427; B.1.429)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Wuhan/HBCDC-HB-04/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Wuhan/HBCDC-HB-04/2020
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Venezuela/Bol285/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Venezuela/Bol285/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/USA/CA-CZB-14678/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/USA/CA-CZB-14678/2020
|
||||||||
| Strains Family |
Epsilon (B.1.427; B.1.429)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Switzerland/TI-EOC-902_25839324/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Switzerland/TI-EOC-902_25839324/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Suriname/SR-342/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Suriname/SR-342/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Scotland/QEUH-13A999F/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Scotland/QEUH-13A999F/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Scotland/EDB14267/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Scotland/EDB14267/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Philippines/PH-PGC-104159/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Philippines/PH-PGC-104159/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Peru/CAL-INS-5458/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Peru/CAL-INS-5458/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/NewZealand/20CV0676/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/NewZealand/20CV0676/2020
|
||||||||
| Strains Family |
Epsilon (B.1.427; B.1.429)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Namibia/N17380/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Namibia/N17380/2021
|
||||||||
| Strains Family |
Delta (B.1.617.2 and AY lineages)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Namibia/CERI-KRISP-K022530/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Namibia/CERI-KRISP-K022530/2021
|
||||||||
| Strains Family |
Delta (B.1.617.2 and AY lineages)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Myanmar/DSMRC-065/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Myanmar/DSMRC-065/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Malaysia/MGI_GS0622/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Malaysia/MGI_GS0622/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Malawi/KRISP-K010154/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Malawi/KRISP-K010154/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Kyrgyzstan/ChVir26576_10/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Kyrgyzstan/ChVir26576_10/2021
|
||||||||
| Strains Family |
Delta (B.1.617.2 and AY lineages)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Kazakhstan/1872/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Kazakhstan/1872/2021
|
||||||||
| Strains Family |
Delta (B.1.617.2 and AY lineages)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Iraq/Thi-Qar-1/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Iraq/Thi-Qar-1/2021
|
||||||||
| Strains Family |
Delta (B.1.617.2 and AY lineages)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Guam/GU-CDC-2-3906081-/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Guam/GU-CDC-2-3906081-/2021
|
||||||||
| Strains Family |
Epsilon (B.1.427; B.1.429)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Ethiopia/ILRI_COVM02835/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Ethiopia/ILRI_COVM02835/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Eswatini/N2779/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Eswatini/N2779/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/QEUH-1269F5E/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/England/QEUH-1269F5E/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/MILK-F88D46/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/England/MILK-F88D46/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/MILK-F88C2B/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/England/MILK-F88C2B/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/MILK-1205929/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/England/MILK-1205929/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/MILK-11F02FA/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/England/MILK-11F02FA/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/LOND-12F444A/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/England/LOND-12F444A/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/CAMC-13B6BB7/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/England/CAMC-13B6BB7/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/England/CAMC-12DF946/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/England/CAMC-12DF946/2021
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/ElSalvador/INC-LNSP-153/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/ElSalvador/INC-LNSP-153/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Chile/AT-59206/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Chile/AT-59206/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Canada/UN-276648/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Canada/UN-276648/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Bonaire/BQ-RIVM-101067/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Bonaire/BQ-RIVM-101067/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Bangladesh/CHRF-GSB17-117/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Bangladesh/CHRF-GSB17-117/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Australia/SA60463/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Australia/SA60463/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Argentina/PAIS-E0546/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Argentina/PAIS-E0546/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Argentina/INEI113521/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Argentina/INEI113521/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Argentina/INEI106123/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Argentina/INEI106123/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Argentina/INEI104133/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/Argentina/INEI104133/2021
|
||||||||
| Strains Family |
Gamma (P.1)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-OC43/VR-759 Quebec/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[7] | |||||||
| Strains Name |
hCov-OC43/VR-759 Quebec/2019
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus OC43 (HCov-OC43)
|
||||||||
| Infection Cells | HCT-8 cells (Human ileocaecal adenocarcinoma cell) HCT-8 cells (Human ileocaecal adenocarcinoma cell) (CVCL_2478 ) | ||||||||
| Cell Originated Tissue | Ileocecum | ||||||||
| Infection Time | 12h; 24; 36h; 48h | ||||||||
| Interaction Score | p_value < 0.05; FDR < 10% | ||||||||
| Method Description | Modified RNA antisense purification coupled with mass spectrometry (RAP-MS); label-free quantification (LFQ); liquid chromatography with tandem mass spectrometry (LC-MS/MS) | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/USA/WA1/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[8] | |||||||
| Strains Name |
hCoV-19/USA/WA1/2020
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7.5 cells (Hepatocyte derived cellular carcinoma cell) Huh7.5 cells (Hepatocyte derived cellular carcinoma cell) (CVCL_7927 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 48 h | ||||||||
| Interaction Score | FDR ≤ 0.05 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[9] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (liver carcinoma cell) Huh7 cells (liver carcinoma cell) (CVCL_0336 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 24h | ||||||||
| Interaction Score | log2FC = 1.19875E+14 | ||||||||
| Method Description | RNA antisense purification and quantitative mass spectrometry (RAP-MS); Tandem mass tag (TMT) labelling; liquid chromatography tandem mass spectrometry (LC-MS/MS); Westernblot | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/England/02/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/England/02/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Calu-3 cells (Human lung cancer cell) Calu-3 cells (Human lung cancer cell) (CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Lung | ||||||||
| Infection Time | 24 h | ||||||||
| Interaction Score | P-adjust = 0.013 | ||||||||
| Method Description | UV protein-RNA crosslinking; RNA interactome capture (cRIC); RNA antisense purification coupled with mass spectrometry (RAP-MS) | ||||||||
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
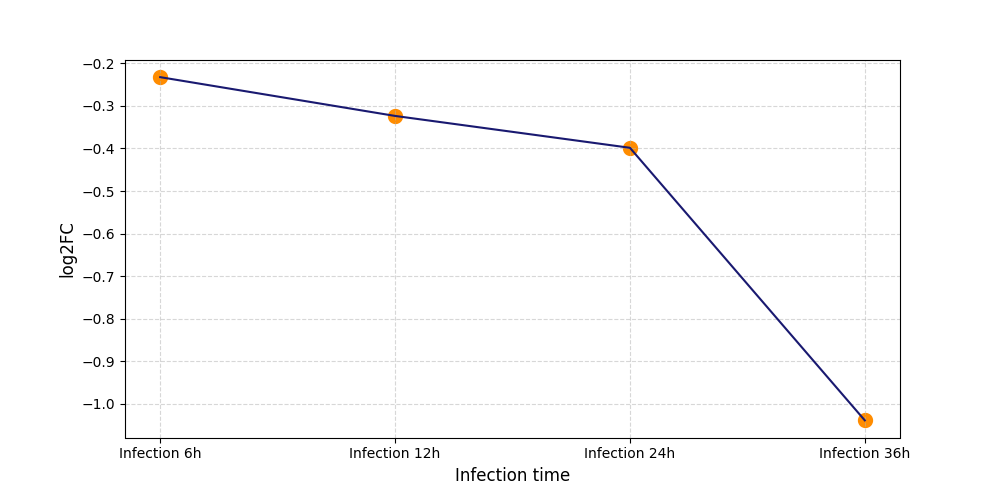
S594
[11] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
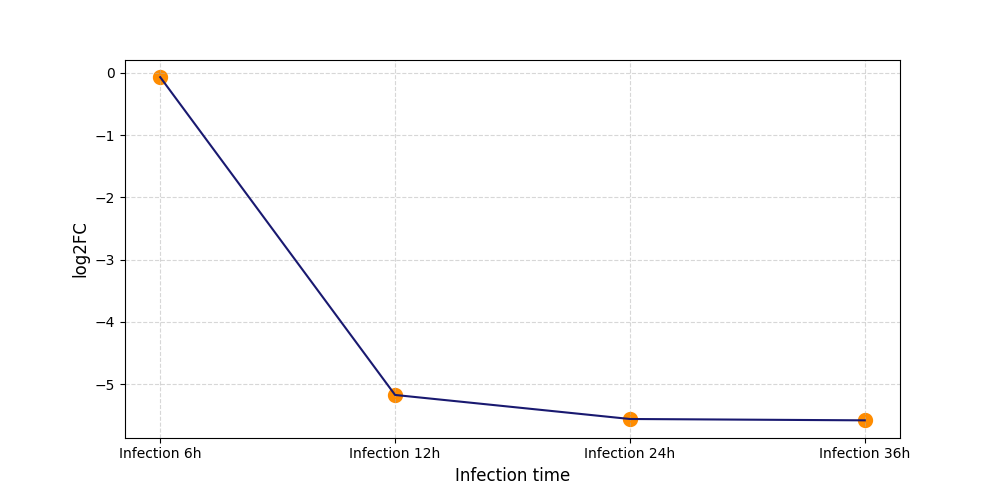
S612
[11] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
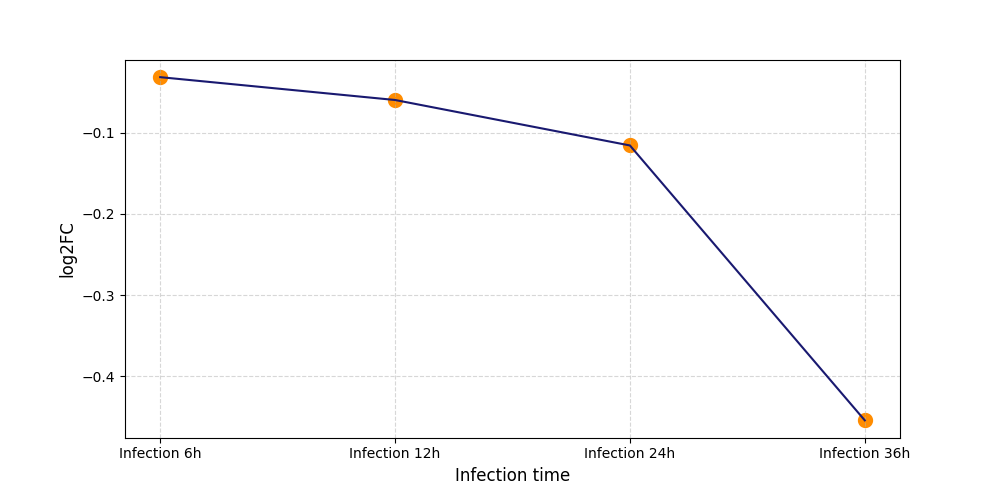
T323
[11] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MSHVAVENALGLDQQFAGLDLNSSDNQSGGSTASKGRYIPPHLRNREATKGFYDKDSSGWSSSKDKDAYSSFGSRSDSRGKSSFFSDRGSGSRGRFDDRGRSDYDGIGSRGDRSGFGKFERGGNSRWCDKSDEDDWSKPLPPSERLEQELFSGGNTGINFEKYDDIPVEATGNNCPPHIESFSDVEMGEIIMGNIELTRYTRPTPVQKHAIPIIKEKRDLMACAQTGSGKTAAFLLPILSQIYSDGPGEALRAMKENGRYGRRKQYPISLVLAPTRELAVQIYEEARKFSYRSRVRPCVVYGGADIGQQIRDLERGCHLLVATPGRLVDMMERGKIGLDFCKYLVLDEADRMLDMGFEPQIRRIVEQDTMPPKGVRHTMMFSATFPKEIQMLARDFLDEYIFLAVGRVGSTSENITQKVVWVEESDKRSFLLDLLNATGKDSLTLVFVETKKGADSLEDFLYHEGYACTSIHGDRSQRDREEALHQFRSGKSPILVATAVAARGLDISNVKHVINFDLPSDIEEYVHRIGRTGRVGNLGLATSFFNERNINITKDLLDLLVEAKQEVPSWLENMAYEHHYKGSSRGRSKSSRFSGGFGARDYRQSSGASSSSFSSSRASSSRSGGGGHGSSRGFGGGGYGGFYNSDGYGGNYNSQGVDWWGN
Click to Show/Hide
|
|---|