Details of Host Protein
| Host Protein General Information (ID: PT0758) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
Polyadenylate-binding protein 1 (PABPC1)
|
Gene Name |
PABPC1
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
PABP1_HUMAN
|
||||||
| Protein Families |
Polyadenylate-binding protein type-1 family
|
||||||||
| Subcellular Location |
Cytoplasm; Stress granule Nucleus Cell projection; lamellipodium
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
Binds the poly (A) tail of mRNA, including that of its owntranscript, and regulates processes of mRNA metabolism such as pre-mRNAsplicing and mRNA stability. Its function in translational initiation regulationcan either be enhanced by PAIP1 or repressed by PAIP2. Can probably bind to cytoplasmic RNA sequences otherthan poly (A) in vivo. Binds to N6-methyladenosine (m6A) -containingmRNAs and contributes to MYC stability by binding to m6A-containing MYCmRNAs. Involved in translationally coupled mRNAturnover. Implicated with other RNA-binding proteinsin the cytoplasmic deadenylation/translational and decay interplay ofthe FOS mRNA mediated by the major coding-region determinant ofinstability (mCRD) domain. Involved in regulation ofnonsense-mediated decay (NMD) of mRNAs containing premature stopcodons; for the recognition of premature termination codons (PTC) andinitiation of NMD a competitive interaction between UPF1 and PABPC1with the ribosome-bound release factors is proposed. By binding to long poly (A) tails, may protect them from uridylation byZCCHC6/ZCCHC11 and hence contribute to mRNA stability.
[1-4]
Click to Show/Hide
|
||||||||
| Related KEGG Pathway | |||||||||
| mRNA surveillance pathway | hsa03015 |
Pathway Map 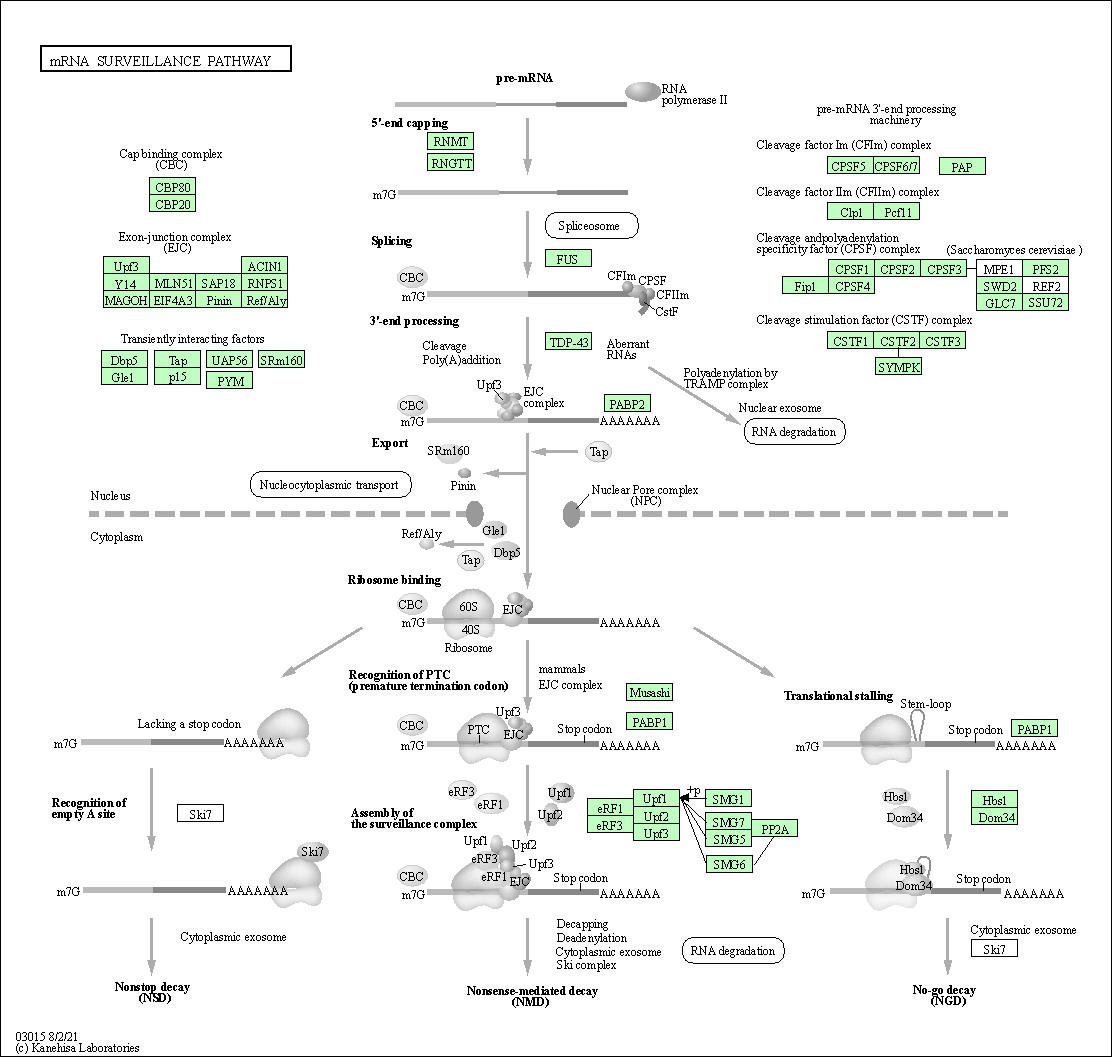
|
|||||||
| RNA degradation | hsa03018 |
Pathway Map 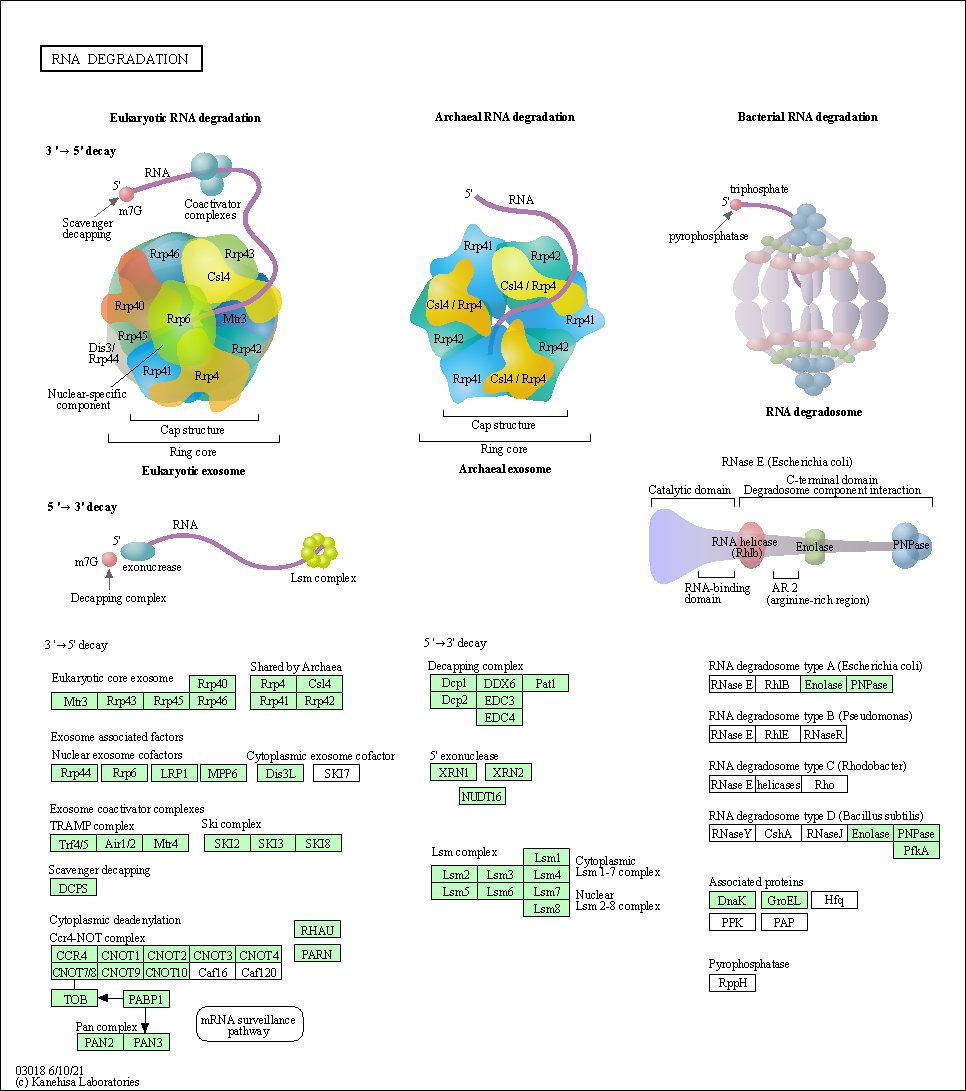
|
|||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Anti-viral | [5] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein PABPC1 in viral infection, PABPC1 protein knockout Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of PABPC1 increases SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: 3'-UTR (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: 3'-UTR (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[7] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) Huh7 cells (human liver cell line); Calu-3 cells (human lung cancer cell line) (CVCL_0336;CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Liver; Lung | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| RNA Region: 3'-UTR (hCoV-19/IPBCAMS-YL01/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[8], [9], [10] | |||||||
| Strains Name |
hCoV-19/IPBCAMS-YL01/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | known to be direct binder | ||||||||
| Interaction Binding Type | Single stranded RNA binding | ||||||||
| Infection Cells | Huh7.5.1 cells (Hepatocyte derived cellular carcinoma cell) Huh7.5.1 cells (Hepatocyte derived cellular carcinoma cell) (CVCL_E049 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 30 h | ||||||||
| Interaction Score | MIST = 0.685537574 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||
| RNA Region: 5'-UTR (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: 5'-UTR (hCoV-19/South Korea/KCDC03/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[11] | |||||||
| Strains Name |
hCoV-19/South Korea/KCDC03/2020
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Directly bind to SARS-CoV-23 RNA's 5' UTR region | ||||||||
| Infection Cells | Vero cells (epithelial kidney cell) Vero cells (epithelial kidney cell) (CVCL_0059 ) | ||||||||
| Cell Originated Tissue | kidney | ||||||||
| Infection Time | 24 h | ||||||||
| Interaction Score | P-adjust < 0.05 | ||||||||
| Method Description | Modified RNA antisense purification coupled with mass spectrometry (RAP-MS); label-free quantification (LFQ); liquid chromatography with tandem mass spectrometry (LC-MS/MS) | ||||||||
| RNA Region: ORF1ab (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF1ab
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF3a (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF3a
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF6 (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF6
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF7a (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF7a
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF7b (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF7b
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: ORF8 (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
ORF8
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: E region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
E region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: M region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
M region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: N region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: S region (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Method Description | ATtRACT Database; Position Weight Matrices (PWMs); TFBSTools R package | ||||||||
| RNA Region: Not Specified Virus Region (hCov-OC43/VR-759 Quebec/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[11] | |||||||
| Strains Name |
hCov-OC43/VR-759 Quebec/2019
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus OC43 (HCov-OC43)
|
||||||||
| Infection Cells | HCT-8 cells (Human ileocaecal adenocarcinoma cell) HCT-8 cells (Human ileocaecal adenocarcinoma cell) (CVCL_2478 ) | ||||||||
| Cell Originated Tissue | Ileocecum | ||||||||
| Infection Time | 12h; 24; 36h; 48h | ||||||||
| Interaction Score | p_value < 0.05; FDR < 10% | ||||||||
| Method Description | Modified RNA antisense purification coupled with mass spectrometry (RAP-MS); label-free quantification (LFQ); liquid chromatography with tandem mass spectrometry (LC-MS/MS) | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/USA/WA1/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[12] | |||||||
| Strains Name |
hCoV-19/USA/WA1/2020
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7.5 cells (Hepatocyte derived cellular carcinoma cell) Huh7.5 cells (Hepatocyte derived cellular carcinoma cell) (CVCL_7927 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 48 h | ||||||||
| Interaction Score | FDR ≤ 0.05 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[13] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (liver carcinoma cell) Huh7 cells (liver carcinoma cell) (CVCL_0336 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 24h | ||||||||
| Interaction Score | log2FC = 2.28233E+14 | ||||||||
| Method Description | RNA antisense purification and quantitative mass spectrometry (RAP-MS); Tandem mass tag (TMT) labelling; liquid chromatography tandem mass spectrometry (LC-MS/MS); Westernblot | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/France/IDF-220-95/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[14] | |||||||
| Strains Name |
hCoV-19/France/IDF-220-95/2020
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Direct interaction | ||||||||
| Infection Cells | HEK293 Cells (Human embryonic kidney cell) HEK293 Cells (Human embryonic kidney cell) (CVCL_0045 ) | ||||||||
| Cell Originated Tissue | Kidney | ||||||||
| Infection Time | 48 h | ||||||||
| Interaction Score | SAINT score ≥ 0.79 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||
| RNA Region: cap- and poly(A) (hCoV-19/England/02/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[15] | |||||||
| Strains Name |
hCoV-19/England/02/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
cap- and poly(A)
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Calu-3 cells (Human lung cancer cell) Calu-3 cells (Human lung cancer cell) (CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Lung | ||||||||
| Infection Time | 24 h | ||||||||
| Interaction Score | P-adjust = 0.012 | ||||||||
| Method Description | UV protein-RNA crosslinking; RNA interactome capture (cRIC); RNA antisense purification coupled with mass spectrometry (RAP-MS) | ||||||||
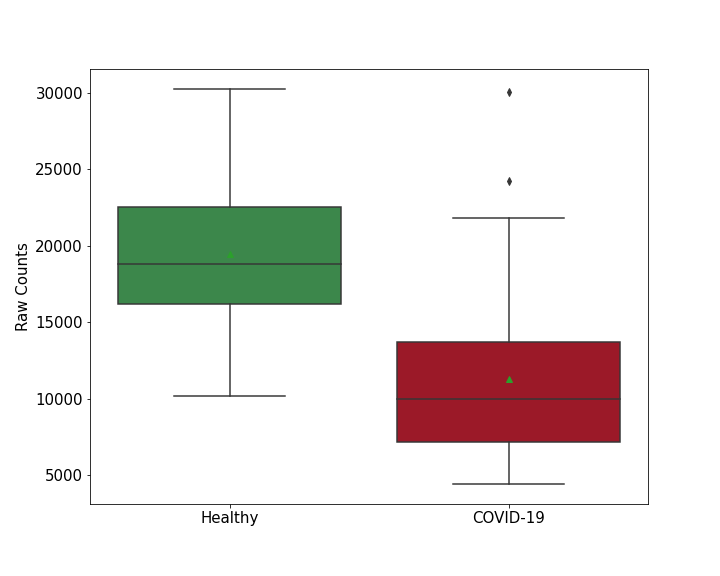
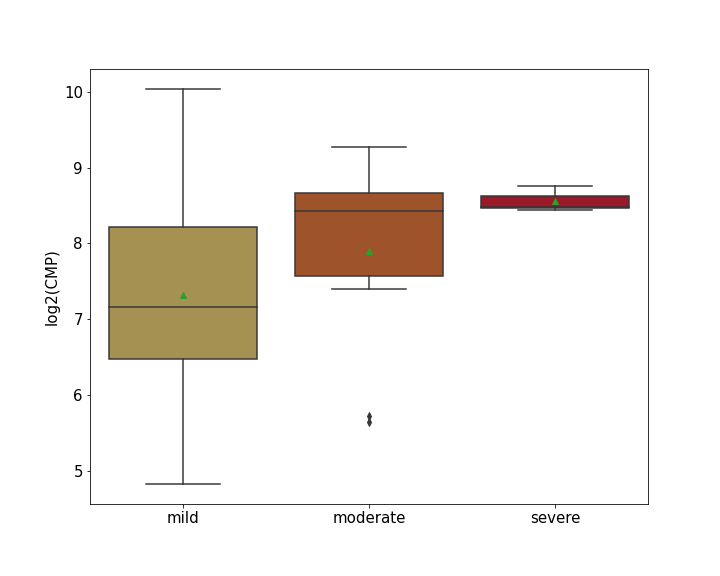
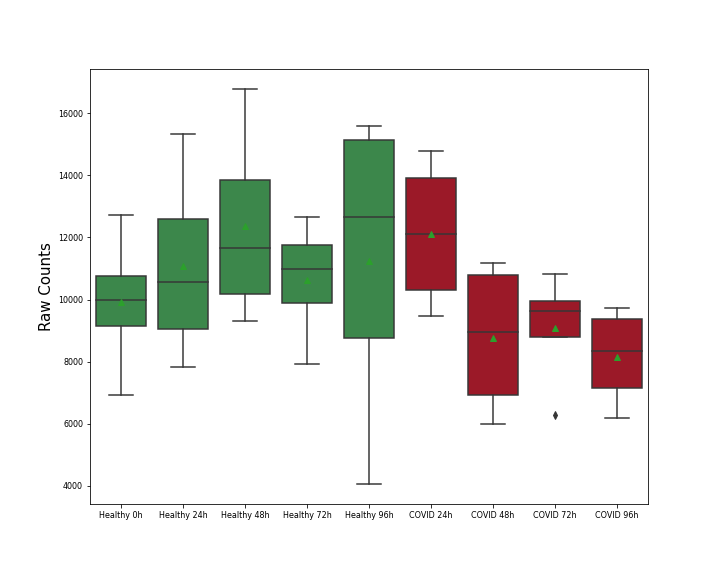
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S470
[16] |
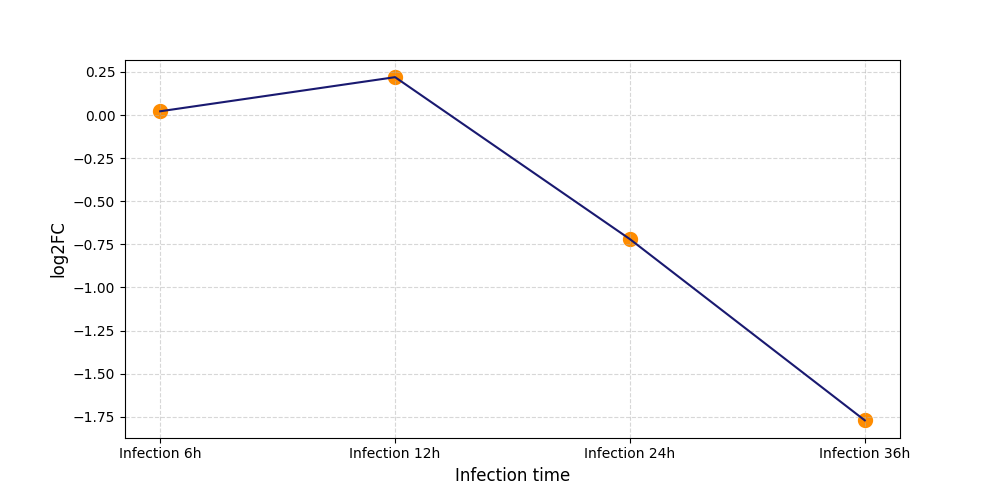 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MNPSAPSYPMASLYVGDLHPDVTEAMLYEKFSPAGPILSIRVCRDMITRRSLGYAYVNFQQPADAERALDTMNFDVIKGKPVRIMWSQRDPSLRKSGVGNIFIKNLDKSIDNKALYDTFSAFGNILSCKVVCDENGSKGYGFVHFETQEAAERAIEKMNGMLLNDRKVFVGRFKSRKEREAELGARAKEFTNVYIKNFGEDMDDERLKDLFGKFGPALSVKVMTDESGKSKGFGFVSFERHEDAQKAVDEMNGKELNGKQIYVGRAQKKVERQTELKRKFEQMKQDRITRYQGVNLYVKNLDDGIDDERLRKEFSPFGTITSAKVMMEGGRSKGFGFVCFSSPEEATKAVTEMNGRIVATKPLYVALAQRKEERQAHLTNQYMQRMASVRAVPNPVINPYQPAPPSGYFMAAIPQTQNRAAYYPPSQIAQLRPSPRWTAQGARPHPFQNMPGAIRPAAPRPPFSTMRPASSQVPRVMSTQRVANTSTQTMGPRPAAAAAAATPAVRTVPQYKYAAGVRNPQQHLNAQPQVTMQQPAVHVQGQEPLTASMLASAPPQEQKQMLGERLFPLIQAMHPTLAGKITGMLLEIDNSELLHMLESPESLRSKVDEAVAVLQAHQAKEAAQKAVNSATGVPTV
Click to Show/Hide
|
|---|