Details of Host Protein
| Host Protein General Information (ID: PT0571) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Name |
IGF2-binding protein 3 (IMP-3)
|
Gene Name |
IGF2BP3
|
||||||
| Host Species |
Homo sapiens
|
Uniprot Entry Name |
IF2B3_HUMAN
|
||||||
| Protein Families |
RRM IMP/VICKZ family
|
||||||||
| Subcellular Location |
Nucleus
|
||||||||
| External Link | |||||||||
| NCBI Gene ID | |||||||||
| Uniprot ID | |||||||||
| Ensembl ID | |||||||||
| HGNC ID | |||||||||
| Function in Host |
RNA-binding factor that may recruit target transcripts tocytoplasmic protein-RNA complexes (mRNPs). This transcript 'caging'into mRNPs allows mRNA transport and transient storage. It alsomodulates the rate and location at which target transcripts encounterthe translational apparatus and shields them from endonuclease attacksor microRNA-mediated degradation. Preferentially binds to N6-methyladenosine (m6A) -containing mRNAs and increases their stability. Binds to the 3'-UTR of CD44 mRNA and stabilizes it, hence promotes cell adhesion and invadopodia formation in cancer cells. Binds to beta-actin/ACTB and MYC transcripts. Increases MYC mRNAstability by binding to the coding region instability determinant (CRD) and binding is enhanced by m6A-modification of the CRD. Binds to the 5'-UTR of the insulin-like growthfactor 2 (IGF2) mRNAs.
[1-2]
Click to Show/Hide
|
||||||||
| 3D Structure |
|
||||||||
| Function of This Protein During Virus Infection | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Name | SARS-COV-2 | Protein Function | Anti-viral | [3] | |||||
| Infected Tissue | Lung | Infection Time | 7-9 Days | ||||||
| Infected Cell | Calu-3 Cells (Human epithelial cell line) | Cellosaurus ID | CVCL_0609 | ||||||
| Method Description | To detect the role of host protein IGF2BP3 in viral infection, IGF2BP3 protein knockout Calu-3 Cells were infected with SARS-COV-2 for 7 - 9 Days , and the effects on infection was detected through CRISPR-based genome-wide gene-knockout screen. | ||||||||
| Results | It is reported that knockout of IGF2BP3 increases SARS-CoV-2 RNA levels compared with control group. | ||||||||
| Host Protein - Virus RNA Network | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Full List of Virus RNA Interacting with This Protien | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| RNA Region: 3'-UTR (hCoV-19/IPBCAMS-YL01/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[4] | |||||||
| Strains Name |
hCoV-19/IPBCAMS-YL01/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7.5.1 cells (Hepatocyte derived cellular carcinoma cell) (CVCL_E049 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 30 h | ||||||||
| Interaction Score | Affinity = 0.996 | ||||||||
| Method Description | PrismNet; pull-down western (STAR methods); Mass spectrometry | ||||||||
| RNA Region: 3'-UTR (hCoV-19/IPBCAMS-YL01/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[5] | |||||||
| Strains Name |
hCoV-19/IPBCAMS-YL01/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
3'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potiential to be direct binder | ||||||||
| Interaction Binding Type | Single stranded RNA binding | ||||||||
| Infection Cells | Huh7.5.1 cells (Hepatocyte derived cellular carcinoma cell) Huh7.5.1 cells (Hepatocyte derived cellular carcinoma cell) (CVCL_E049 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 30 h | ||||||||
| Interaction Score | MIST = 0.685511996 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||
| RNA Region: 5'-UTR (hCoV-SARS/Manitoba/2003 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-SARS/Manitoba/2003
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome-related coronavirus (HCoV-SARS)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 5'-UTR (hCov-OC43/VR-759 Quebec/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-OC43/VR-759 Quebec/2019
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Human coronavirus OC43 (HCov-OC43)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 5'-UTR (hCov-NL63/Amsterdam/2004 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-NL63/Amsterdam/2004
|
||||||||
| Strains Family |
Alpha
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Human Coronavirus NL63 (HCov-NL63)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 5'-UTR (hCov-MERS/Jeddah/2012 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-MERS/Jeddah/2012
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Middle East respiratory syndrome-related coronavirus (HCov-MERS)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 5'-UTR (hCoV-HKU1/Hong Kong/2004 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-HKU1/Hong Kong/2004
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Human coronavirus HKU1 (HCoV-HKU1)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 5'-UTR (hCov-229E/Würzburg/2000 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCov-229E/Würzburg/2000
|
||||||||
| Strains Family |
Alpha
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Human coronavirus 229E (HCov-229E)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 5'-UTR (hCoV-19/Wuhan-Hu-1/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan-Hu-1/2019
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: 5'-UTR (hCoV-19/South Korea/KCDC03/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[7] | |||||||
| Strains Name |
hCoV-19/South Korea/KCDC03/2020
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Directly bind to SARS-CoV-23 RNA's 5' UTR region | ||||||||
| Infection Cells | Vero cells (epithelial kidney cell) Vero cells (epithelial kidney cell) (CVCL_0059 ) | ||||||||
| Cell Originated Tissue | kidney | ||||||||
| Infection Time | 24 h | ||||||||
| Interaction Score | P-adjust < 0.05 | ||||||||
| Method Description | Modified RNA antisense purification coupled with mass spectrometry (RAP-MS); label-free quantification (LFQ); liquid chromatography with tandem mass spectrometry (LC-MS/MS) | ||||||||
| RNA Region: 5'-UTR (hCoV-19/IPBCAMS-YL01/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[4] | |||||||
| Strains Name |
hCoV-19/IPBCAMS-YL01/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
5'-UTR
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7.5.1 cells (Hepatocyte derived cellular carcinoma cell) Huh7.5.1 cells (Hepatocyte derived cellular carcinoma cell) (CVCL_E049 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 30 h | ||||||||
| Interaction Score | Affinity = 0.941 | ||||||||
| Method Description | PrismNet; pull-down western (STAR methods); Mass spectrometry | ||||||||
| RNA Region: ORF10 (hCoV-19/Not Specified Virus Strain ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[8] | |||||||
| Strains Name |
hCoV-19/Not Specified Virus Strain
|
||||||||
| RNA Binding Region |
ORF10
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7 cells (human liver cell line) Huh7 cells (human liver cell line) (CVCL_0336 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Interaction Score | P-value < 0.05 | ||||||||
| Method Description | RNA pull-down assays; liquid chromatography with tandem mass spectrometry (LC-MS/MS); Wilcoxon test; MS2 affinity purification coupled with liquid chromatography-mass spectrometry (MAMS) | ||||||||
| RNA Region: N region (hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Wuhan/HBCDC-HB-04/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan/HBCDC-HB-04/2020
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Philippines/PH-PGC-104159/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Philippines/PH-PGC-104159/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Myanmar/DSMRC-065/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Myanmar/DSMRC-065/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Malaysia/MGI_GS0622/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Malaysia/MGI_GS0622/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Ethiopia/ILRI_COVM02835/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Ethiopia/ILRI_COVM02835/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/ElSalvador/INC-LNSP-153/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/ElSalvador/INC-LNSP-153/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Chile/AT-59206/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Chile/AT-59206/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Canada/UN-276648/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Canada/UN-276648/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Bonaire/BQ-RIVM-101067/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Bonaire/BQ-RIVM-101067/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Bangladesh/CHRF-GSB17-117/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Bangladesh/CHRF-GSB17-117/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: N region (hCoV-19/Australia/SA60463/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Australia/SA60463/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
N region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Zimbabwe/CERI-KRISP-K034087/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/Wuhan/HBCDC-HB-04/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Wuhan/HBCDC-HB-04/2020
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/Philippines/PH-PGC-104159/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Philippines/PH-PGC-104159/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/Myanmar/DSMRC-065/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Myanmar/DSMRC-065/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/Malaysia/MGI_GS0622/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Malaysia/MGI_GS0622/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/Ethiopia/ILRI_COVM02835/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Ethiopia/ILRI_COVM02835/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/ElSalvador/INC-LNSP-153/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/ElSalvador/INC-LNSP-153/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/Chile/AT-59206/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Chile/AT-59206/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/Canada/UN-276648/2021 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Canada/UN-276648/2021
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/Bonaire/BQ-RIVM-101067/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Bonaire/BQ-RIVM-101067/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/Bangladesh/CHRF-GSB17-117/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Bangladesh/CHRF-GSB17-117/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: S region (hCoV-19/Australia/SA60463/2022 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[6] | |||||||
| Strains Name |
hCoV-19/Australia/SA60463/2022
|
||||||||
| Strains Family |
Omicron (B.1.1.529; BA.1; BA.1.1; BA.2; BA.3; BA.4 ; BA.5)
|
||||||||
| RNA Binding Region |
S region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Potential binding protein | ||||||||
| Infection Cells | HepG2 cells (hepatocellular carcinoma cell line) HepG2 cells (hepatocellular carcinoma cell line) (CVCL_0027 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Method Description | ENCODE project database; Pysster model; DeepRiPe (Section 3.6; 3.7; 3.8; 3.9); ClustalO (72) algorithm | ||||||||
| RNA Region: Not Specified Virus Region (hCov-OC43/VR-759 Quebec/2019 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[7] | |||||||
| Strains Name |
hCov-OC43/VR-759 Quebec/2019
|
||||||||
| Strains Family |
Beta
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Human coronavirus OC43 (HCov-OC43)
|
||||||||
| Infection Cells | HCT-8 cells (Human ileocaecal adenocarcinoma cell) HCT-8 cells (Human ileocaecal adenocarcinoma cell) (CVCL_2478 ) | ||||||||
| Cell Originated Tissue | Ileocecum | ||||||||
| Infection Time | 12h; 24; 36h; 48h | ||||||||
| Interaction Score | p_value < 0.05; FDR < 10% | ||||||||
| Method Description | Modified RNA antisense purification coupled with mass spectrometry (RAP-MS); label-free quantification (LFQ); liquid chromatography with tandem mass spectrometry (LC-MS/MS) | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/USA/WA1/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[9] | |||||||
| Strains Name |
hCoV-19/USA/WA1/2020
|
||||||||
| Strains Family |
Alpha (B.1.1.7)
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Huh7.5 cells (Hepatocyte derived cellular carcinoma cell) Huh7.5 cells (Hepatocyte derived cellular carcinoma cell) (CVCL_7927 ) | ||||||||
| Cell Originated Tissue | Liver | ||||||||
| Infection Time | 48 h | ||||||||
| Interaction Score | FDR ≤ 0.05 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/France/IDF-220-95/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[10] | |||||||
| Strains Name |
hCoV-19/France/IDF-220-95/2020
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Interaction Type | Direct interaction | ||||||||
| Infection Cells | HEK293 Cells (Human embryonic kidney cell) HEK293 Cells (Human embryonic kidney cell) (CVCL_0045 ) | ||||||||
| Cell Originated Tissue | Kidney | ||||||||
| Infection Time | 48 h | ||||||||
| Interaction Score | SAINT score ≥ 0.79 | ||||||||
| Method Description | comprehensive identification of RNA-binding proteins by massspectrometry (ChIRP-MS) | ||||||||
| RNA Region: Not Specified Virus Region (hCoV-19/England/02/2020 ) | |||||||||
| RNA Region Details |
RNA Info
 Click to show the detail information of this RNA binding region
Click to show the detail information of this RNA binding region
|
[11] | |||||||
| Strains Name |
hCoV-19/England/02/2020
|
||||||||
| Strains Family |
Beta (B.1.351)
|
||||||||
| RNA Binding Region |
Not Specified Virus Region
|
||||||||
| Virus Name |
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
|
||||||||
| Infection Cells | Calu-3 cells (Human lung cancer cell) Calu-3 cells (Human lung cancer cell) (CVCL_0609 ) | ||||||||
| Cell Originated Tissue | Lung | ||||||||
| Infection Time | 24 h | ||||||||
| Interaction Score | P-adjust = 0.017 | ||||||||
| Method Description | UV protein-RNA crosslinking; RNA interactome capture (cRIC); RNA antisense purification coupled with mass spectrometry (RAP-MS) | ||||||||
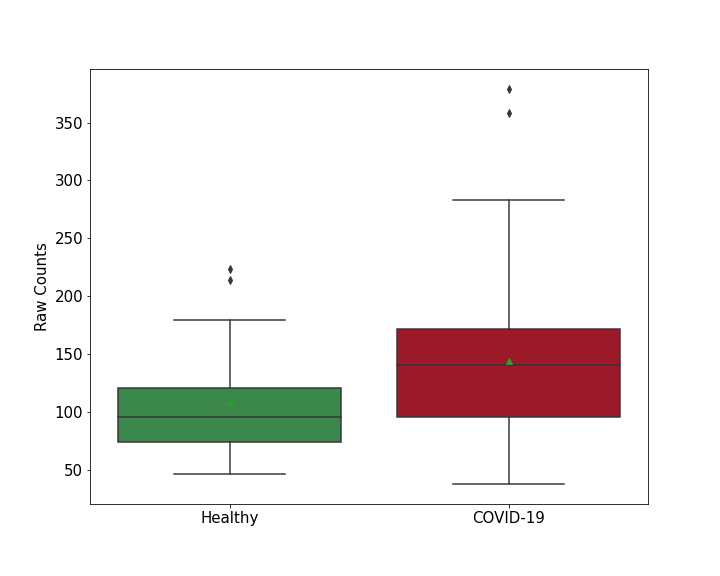
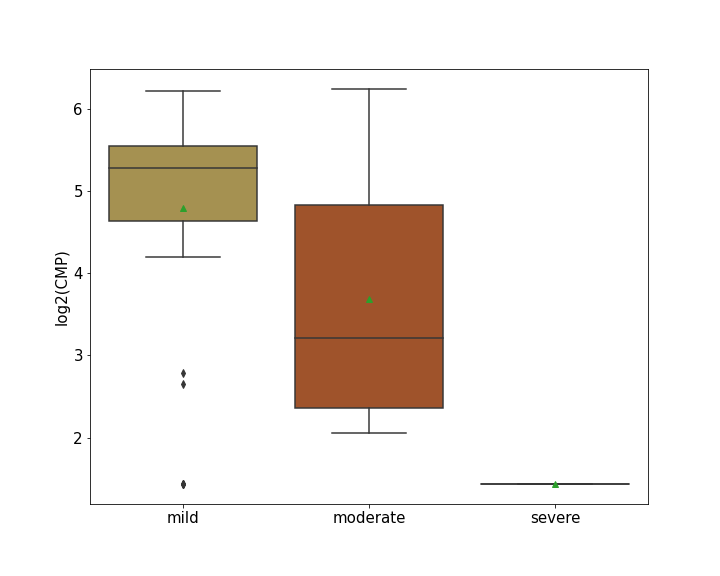
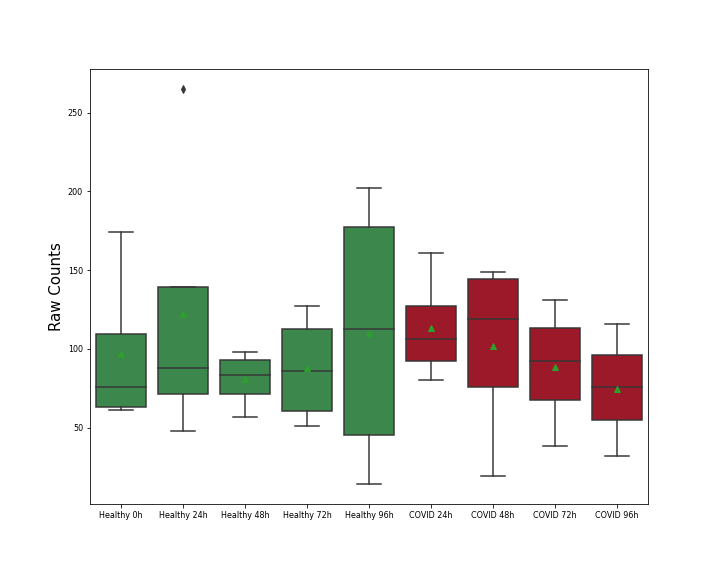
| Differential Gene Expression During SARS-COV-2 Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
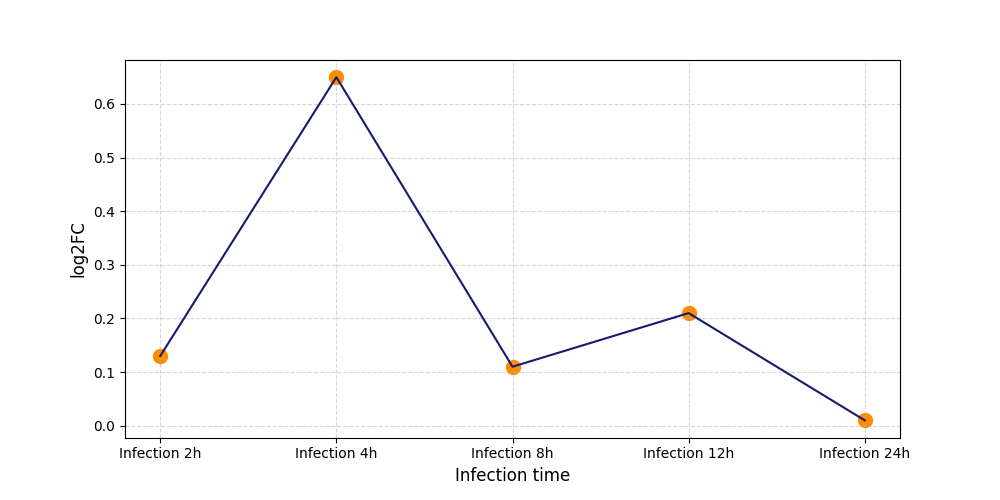
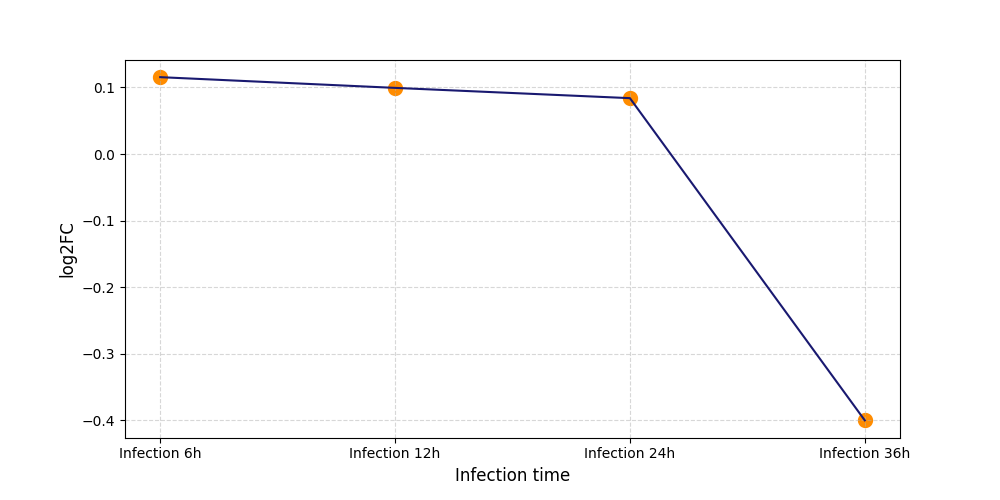
| Protein Phosphorylation after Virus Infection | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
S184
[12] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
S184
[13] |
 |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Sequence Information |
MNKLYIGNLSENAAPSDLESIFKDAKIPVSGPFLVKTGYAFVDCPDESWALKAIEALSGKIELHGKPIEVEHSVPKRQRIRKLQIRNIPPHLQWEVLDSLLVQYGVVESCEQVNTDSETAVVNVTYSSKDQARQALDKLNGFQLENFTLKVAYIPDEMAAQQNPLQQPRGRRGLGQRGSSRQGSPGSVSKQKPCDLPLRLLVPTQFVGAIIGKEGATIRNITKQTQSKIDVHRKENAGAAEKSITILSTPEGTSAACKSILEIMHKEAQDIKFTEEIPLKILAHNNFVGRLIGKEGRNLKKIEQDTDTKITISPLQELTLYNPERTITVKGNVETCAKAEEEIMKKIRESYENDIASMNLQAHLIPGLNLNALGLFPPTSGMPPPTSGPPSAMTPPYPQFEQSETETVHLFIPALSVGAIIGKQGQHIKQLSRFAGASIKIAPAEAPDAKVRMVIITGPPEAQFKAQGRIYGKIKEENFVSPKEEVKLEAHIRVPSFAAGRVIGKGGKTVNELQNLSSAEVVVPRDQTPDENDQVVVKITGHFYACQVAQRKIQEILTQVKQHQQQKALQSGPPQSRRK
Click to Show/Hide
|
|---|